Code
install.packages("dplyr")
install.packages("ggplot2")
install.packages("tidyr")
install.packages("flextable")
install.packages("readxl")
install.packages("here")
install.packages("checkdown") Martin Schweinberger
2026


This tutorial introduces R and RStudio — the programming language and development environment used throughout LADAL. It is aimed at complete beginners with no prior programming experience, and walks through everything you need to get up and running: installing software, understanding the RStudio interface, setting up a reproducible project, and working with R for the first time.
R is a free, open-source programming language designed specifically for data analysis and statistics. It is the most widely used tool for quantitative research in linguistics, the social sciences, and the digital humanities — and for good reason. R gives you complete control over your analysis, produces publication-quality graphics, and keeps your work fully transparent and reproducible.
This tutorial will not turn you into an expert. Its goal is to give you a solid, well-structured foundation: to know where things are, how to think about R, and how to start doing real things with data. The rest of LADAL’s tutorials build from here.
This tutorial has no prerequisites — it is designed for complete beginners. However, the following background tutorials are helpful companions:
ggplot2Martin Schweinberger. 2026. Getting Started with R and RStudio. The Language Technology and Data Analysis Laboratory (LADAL), The University of Queensland, Australia. url: https://ladal.edu.au/tutorials/intror/intror.html (Version 2026.03.27), doi: .
Before diving in, it is worth briefly explaining why R is worth learning.
R is free and open-source — there are no licensing costs, ever. It is the dominant tool for statistical analysis in linguistics, psychology, and the social sciences. It has a vast ecosystem of over 20,000 contributed packages that extend its capabilities to cover almost any analytical task imaginable. Its reproducibility features — the ability to combine code, output, and prose in a single document — mean your analyses can be fully transparent and re-run by anyone. And its visualisation capabilities, particularly through ggplot2, are unmatched.
The learning curve is real but manageable. This tutorial gives you the foundation you need.
Install the packages used in this tutorial (only needed once):
Load the packages at the start of each session:
What you’ll learn: How to install R and RStudio on your computer
Why it matters: You need both installed to follow any LADAL tutorial
Time: ~15–30 minutes (mostly waiting for downloads)
R and RStudio are two separate pieces of software that work together. Think of R as the engine and RStudio as the car — you need both, and you interact almost exclusively with RStudio.
R must be installed before RStudio. Visit cran.r-project.org and select the download for your operating system:
Run the downloaded installer and accept the default settings throughout.
R releases a new version approximately once a year. To check your current version, run R.version$version.string in the console. To update on Windows, the installr package automates the process:
On Mac, download the new version from CRAN and install over the existing version.
Visit posit.co/download/rstudio-desktop and download the free RStudio Desktop version for your operating system. Run the installer and accept the defaults.
After installation, open RStudio (not R directly). RStudio will automatically detect your R installation.
What you’ll learn: How to navigate the four panes of RStudio and what each one does
Key concept: The difference between the Console (run immediately) and the Script Editor (save and reuse)
When you first open RStudio, you will see an interface divided into panes. The screenshot below shows a typical RStudio session with all four panes visible.
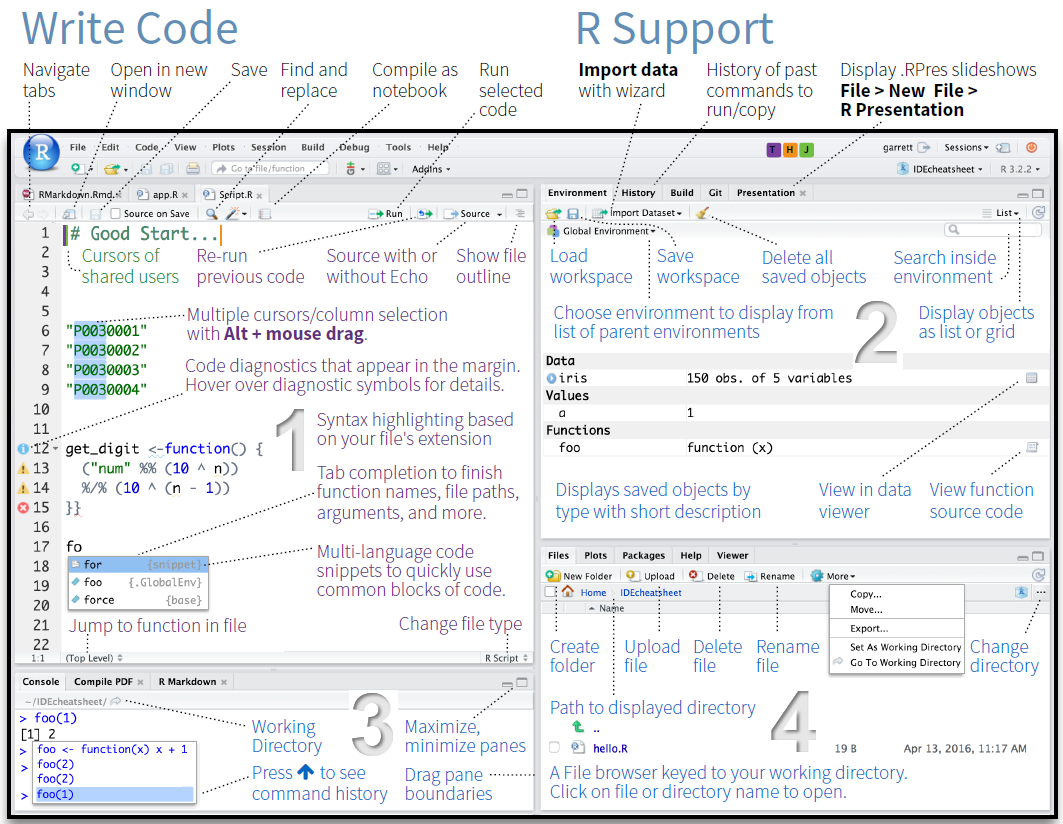
RStudio has four main panes:
This is where you write and save code. Code typed here does not run automatically — you must explicitly execute it. This is where all your analysis lives.
To run a line of code from the Script Editor, place your cursor on that line and press Ctrl + Enter (Windows/Linux) or Cmd + Enter (Mac). To run a highlighted block, select the code first and then press the same shortcut.
This is where R executes code and displays text output. When you run code from the Script Editor, it appears here. You can also type directly into the Console and press Enter to run commands immediately.
Use the Console for quick experiments. Use the Script Editor for anything you want to keep.
Tab to autocomplete?function_name to open the help page for any functionThe Environment tab shows all objects currently loaded in your R session — data frames, variables, vectors, and so on. Clicking on a data frame here opens a spreadsheet-style viewer.
The History tab logs all commands you have run in the current session.
This multi-tab pane contains:
?)What you’ll learn: How to set up a reproducible project in RStudio; what an R Notebook is and why to use one
Key concept: An R Project keeps all your files, code, and data together in one self-contained folder
Good organisation before you start coding saves a great deal of trouble later. This section walks through the recommended setup.
Before opening RStudio, create a folder on your computer for your project. Inside it, create the following sub-folders:
my_project/
├── data/ ← raw and processed data files
├── images/ ← figures saved from R
├── tables/ ← tables exported from R
└── docs/ ← notes, reports, and output documents 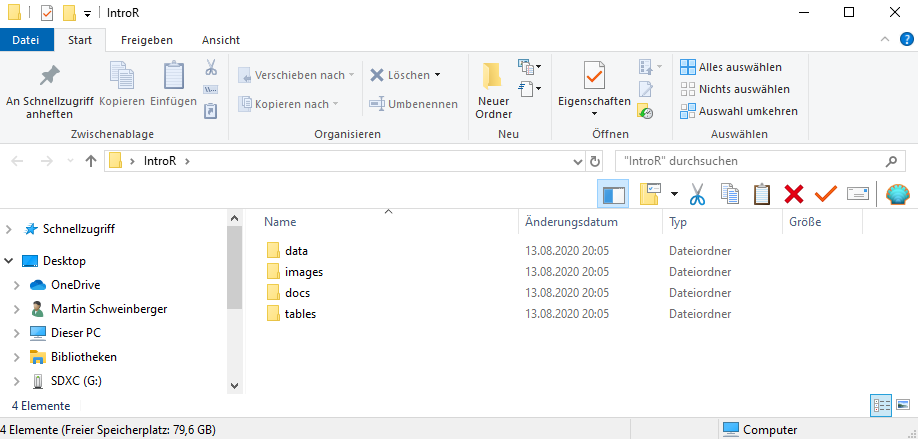
An R Project tells RStudio that a folder is a self-contained project. It sets the working directory automatically (so file paths are predictable) and keeps your project’s history and settings separate from other projects.
To create an R Project:
File → New ProjectExisting DirectoryCreate ProjectRStudio will restart and you will see your project name in the top-right corner. You are now working inside your project.
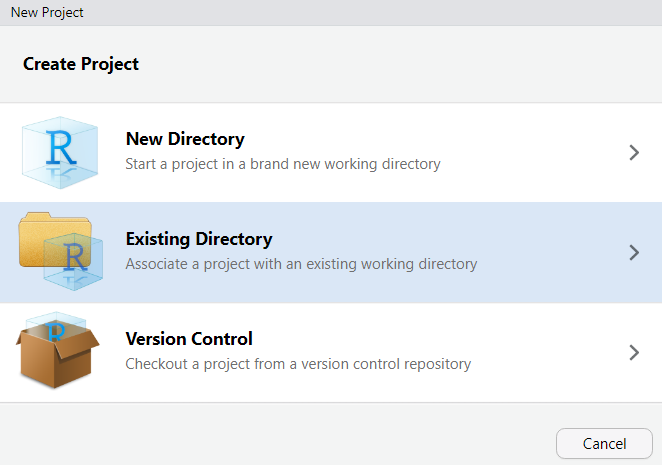
When you open RStudio, always open your project first (either by double-clicking the .Rproj file in your folder, or via File → Open Project). This ensures file paths work correctly and your environment is isolated.
An R Notebook (.Rmd or .qmd file) combines prose, code, and output in a single document. This is the standard format for LADAL tutorials and is highly recommended for your own analyses — it keeps your thinking and your code together.
To create an R Notebook:
File → New File → R Notebook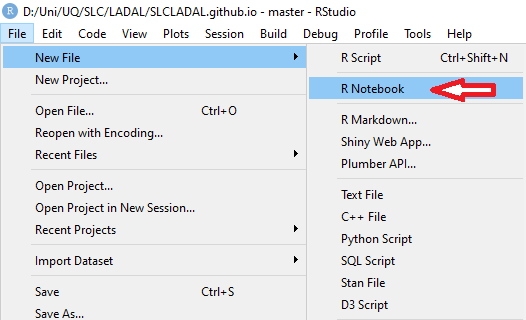
The notebook uses R Markdown — a simple formatting syntax explained below.
R Markdown lets you write formatted prose alongside executable code. Here is a quick reference:
# Heading 1
## Heading 2
### Heading 3
**bold text**
*italic text*
`inline code`
- bullet point
- another bullet
1. numbered item
2. another item
[link text](https://url.com) Code is written inside code chunks (fenced with triple backticks):
::: {.cell}
```{.r .cell-code}
# your R code here
2 + 2
```
::: {.cell-output .cell-output-stdout}
```
[1] 4
```
:::
:::When you click Knit (or Render in Quarto), R Markdown executes all code chunks and weaves the output together with your prose into a finished HTML, PDF, or Word document.
The power of R Notebooks is reproducibility: your entire analysis — every number, table, and figure — is regenerated from scratch each time you render the document. Anyone with your .Rmd file and data can reproduce your results exactly.
What you’ll learn: The core building blocks of R — objects, functions, operators, and assignment
Key concepts: Everything in R is an object; everything you do in R uses a function
At the top of any script or notebook, set global options and load packages. This makes your session reproducible from the very first line.
In R, everything is stored as an object. You create objects using the assignment operator <-:
[1] 42[1] "linguistics"[1] TRUEGood object names are:
- lowercase with underscores for spaces: word_count, not Word Count
- descriptive: reaction_time_ms is better than x
- not starting with a number: data1 is valid; 1data is not
- not reserved words: don’t use c, t, df, mean, TRUE, FALSE, NULL as object names
R is case-sensitive: MyData and mydata are different objects.
A function takes one or more inputs (called arguments), does something, and returns an output. Functions are called by name followed by parentheses containing the arguments:
[1] 12[1] 3.14[1] 11[1] "language-data-analysis"You can nest functions — the inner function runs first:
R provides standard arithmetic and logical operators:
[1] 13[1] 7[1] 30[1] 3.333333[1] 100[1] 1[1] TRUE[1] FALSE[1] TRUE[1] TRUE[1] TRUE[1] FALSE[1] TRUE[1] FALSE= vs ==
One of the most common beginner errors: = is used for assignment (interchangeable with <- in most cases, though <- is preferred); == tests whether two things are equal. 5 = 3 will produce an error; 5 == 3 returns FALSE.
Q1. What does the assignment operator <- do?
Q2. You run my_var <- 10. What will my_var * 3 + 1 return?
Q3. Which of the following is NOT a valid object name in R?
What you’ll learn: The six basic data types in R and why they matter
Key concept: The type of your data determines which operations are valid
Every object in R has a type (also called a class). The four types you will encounter most often are:
[1] "numeric"[1] "integer"[1] "character"[1] "logical"You can check the type of any object with class() or typeof(), and test for specific types:
[1] TRUE[1] TRUE[1] TRUEYou can convert between types using coercion functions:
[1] 3.14[1] "42"[1] FALSE[1] TRUE[1] TRUEWhen R cannot coerce a value, it introduces NA (missing value) with a warning:
Warning: NAs introduced by coercion[1] NANA stands for Not Available and represents missing data. It propagates through calculations — any arithmetic involving NA returns NA unless specifically handled.
What you’ll learn: How R organises collections of data — vectors, data frames, lists, and factors
Key concept: Vectors are the fundamental unit; data frames are collections of equal-length vectors
A vector is a sequence of values of the same type. Vectors are created with c() (short for combine):
You can perform operations on entire vectors at once — R applies them element-by-element:
[1] 6 10 4 16 8 12 2[1] FALSE TRUE FALSE TRUE FALSE TRUE FALSE[1] 7[1] 29[1] 4.142857[1] 2.410295[1] 1[1] 8[1] 1 8 [1] 1 2 3 4 5 6 7 8 9 10[1] 0.00 0.25 0.50 0.75 1.00[1] 1.00 25.75 50.50 75.25 100.00[1] "yes" "yes" "yes"[1] "A" "B" "A" "B" "A" "B" "A" "B"[1] "A" "A" "A" "A" "B" "B" "B" "B"A factor is a special type of vector for categorical variables. Factors have a fixed set of levels (categories) and are essential for grouping in analyses and plots.
[1] Formal Informal Formal ReadAloud Informal
Levels: Formal Informal ReadAloud[1] "Formal" "Informal" "ReadAloud"[1] 3register
Formal Informal ReadAloud
2 2 1 By default, levels are ordered alphabetically. You can specify a custom order:
A data frame is R’s equivalent of a spreadsheet — a table where each column is a vector of the same length. Data frames are the most common way to store linguistic data.
# Create a data frame from scratch
speakers <- data.frame(
ID = 1:6,
Name = c("Alice", "Bob", "Carol", "David", "Eve", "Frank"),
L1 = c("English", "German", "English", "Mandarin", "English", "Arabic"),
Age = c(24, 31, 28, 22, 35, 27),
Proficiency = factor(c("Advanced", "Intermediate", "Advanced",
"Beginner", "Intermediate", "Advanced"),
levels = c("Beginner", "Intermediate", "Advanced"))
)
# Inspect the data frame
speakers ID Name L1 Age Proficiency
1 1 Alice English 24 Advanced
2 2 Bob German 31 Intermediate
3 3 Carol English 28 Advanced
4 4 David Mandarin 22 Beginner
5 5 Eve English 35 Intermediate
6 6 Frank Arabic 27 AdvancedKey functions for inspecting a data frame:
[1] 6[1] 5[1] 6 5[1] "ID" "Name" "L1" "Age" "Proficiency"'data.frame': 6 obs. of 5 variables:
$ ID : int 1 2 3 4 5 6
$ Name : chr "Alice" "Bob" "Carol" "David" ...
$ L1 : chr "English" "German" "English" "Mandarin" ...
$ Age : num 24 31 28 22 35 27
$ Proficiency: Factor w/ 3 levels "Beginner","Intermediate",..: 3 2 3 1 2 3 ID Name L1 Age Proficiency
1 1 Alice English 24 Advanced
2 2 Bob German 31 Intermediate
3 3 Carol English 28 Advanced ID Name L1 Age Proficiency
5 5 Eve English 35 Intermediate
6 6 Frank Arabic 27 Advanced ID Name L1 Age
Min. :1.00 Length:6 Length:6 Min. :22.00
1st Qu.:2.25 Class :character Class :character 1st Qu.:24.75
Median :3.50 Mode :character Mode :character Median :27.50
Mean :3.50 Mean :27.83
3rd Qu.:4.75 3rd Qu.:30.25
Max. :6.00 Max. :35.00
Proficiency
Beginner :1
Intermediate:2
Advanced :3
A list is the most flexible data structure — it can hold objects of different types and lengths, including other lists.
[1] "Study 1"[1] 30Lists are commonly returned by statistical model functions (e.g., lm() returns a list). You rarely create them from scratch but frequently need to extract elements from them.
Q1. You run x <- c(1, 2, "three", 4). What type will x be?
Q2. What is the difference between a factor and a character vector?
Q3. What does dim(df) return for a data frame with 50 rows and 4 columns?
What you’ll learn: How to access specific elements, rows, columns, and subsets of your data
Key concept: Square brackets [ ] select by position; $ selects columns by name; dplyr verbs filter by condition
Extracting exactly the data you need is one of the most fundamental R skills.
Use square brackets [ ] with a position number (index) to extract elements from a vector. R indexing starts at 1 (not 0 as in Python).
[1] "English"[1] "Arabic"[1] "English" "Mandarin"[1] "German" "Mandarin" "Arabic" [1] "English" "Mandarin" "Arabic" "French" [1] "German" "Mandarin" "Arabic" [1] 5 8 6[1] 1Data frames have two dimensions: df[row, column]. Leave one blank to select all rows or all columns.
[1] "German" ID Name L1 Age Proficiency
1 1 Alice English 24 Advanced[1] "English" "German" "English" "Mandarin" "English" "Arabic" [1] 24 31 28 22 35 27[1] "English" "German" "English" "Mandarin" "English" "Arabic" Name Age
1 Alice 24
2 Bob 31
3 Carol 28dplyrWhile base R indexing works, the dplyr package provides cleaner, more readable syntax for filtering and selecting data. These are the two most important dplyr verbs for subsetting:
ID Name L1 Age Proficiency
1 1 Alice English 24 Advanced
2 3 Carol English 28 Advanced
3 5 Eve English 35 Intermediate Name Age Proficiency
1 Alice 24 Advanced
2 Bob 31 Intermediate
3 Carol 28 Advanced
4 David 22 Beginner
5 Eve 35 Intermediate
6 Frank 27 Advanced Name L1 Age
1 Alice English 24
2 Carol English 28
3 David Mandarin 22
4 Frank Arabic 27|>
The pipe |> (from the magrittr/dplyr packages) passes the result on the left to the function on the right. It lets you chain operations in a readable left-to-right sequence instead of nesting functions:
# Without pipe (hard to read)
select(filter(speakers, Age < 30), Name, Age)
# With pipe (reads like a sentence)
speakers |> filter(Age < 30) |> select(Name, Age) R 4.1+ also has a native pipe |> that works similarly. LADAL tutorials use |>.
Q1. Given v <- c(10, 20, 30, 40, 50), what does v[c(2, 4)] return?
Q2. How do you use dplyr::filter() to keep only rows where the column Proficiency equals "Advanced"?
What you’ll learn: How to load data from files, inspect it, and perform common data manipulation operations
Key functions: read.csv(), readxl::read_excel(), dplyr::mutate(), dplyr::group_by(), dplyr::summarise()
# Save as CSV
write.csv(my_data, here::here("data", "processed_data.csv"), row.names = FALSE)
# Save as R object (preserves factors and other R-specific attributes)
saveRDS(my_data, here::here("data", "processed_data.rds"))
# Load an RDS file
my_data <- readRDS(here::here("data", "processed_data.rds")) We will use a simulated linguistic dataset to demonstrate the key dplyr operations. The dataset contains reaction times and accuracy from a lexical decision task:
set.seed(42)
n <- 60
lex_data <- data.frame(
Participant = rep(1:20, each = 3),
Condition = rep(c("High_Freq", "Low_Freq", "Pseudoword"), times = 20),
RT_ms = c(
rnorm(20, mean = 480, sd = 55), # High frequency: fast
rnorm(20, mean = 610, sd = 70), # Low frequency: slower
rnorm(20, mean = 730, sd = 80) # Pseudowords: slowest
),
Accurate = sample(c(TRUE, FALSE), n, replace = TRUE, prob = c(0.9, 0.1))
) |>
dplyr::mutate(Condition = factor(Condition,
levels = c("High_Freq", "Low_Freq", "Pseudoword"))) mutate() — Add or Modify Columns Participant Condition RT_ms Accurate RT_s RT_log Fast_respons
1 1 High_Freq 555.4027 TRUE 0.5554027 6.319693 FALSE
2 1 Low_Freq 448.9416 TRUE 0.4489416 6.106893 TRUE
3 1 Pseudoword 499.9721 TRUE 0.4999721 6.214552 TRUE
4 2 High_Freq 514.8074 TRUE 0.5148074 6.243793 FALSE
5 2 Low_Freq 502.2348 TRUE 0.5022348 6.219068 FALSE
6 2 Pseudoword 474.1632 TRUE 0.4741632 6.161551 TRUEgroup_by() and summarise() — Aggregate by Grouplex_data |>
dplyr::group_by(Condition) |>
dplyr::summarise(
n = n(),
M_RT = round(mean(RT_ms), 1),
SD_RT = round(sd(RT_ms), 1),
Accuracy = round(mean(Accurate) * 100, 1),
.groups = "drop"
) |>
flextable() |>
flextable::set_table_properties(width = .8, layout = "autofit") |>
flextable::theme_zebra() |>
flextable::fontsize(size = 12) |>
flextable::fontsize(size = 12, part = "header") |>
flextable::align_text_col(align = "center") |>
flextable::set_caption(caption = "Reaction times and accuracy by condition in the lexical decision task.") |>
flextable::border_outer() Condition | n | M_RT | SD_RT | Accuracy |
|---|---|---|---|---|
High_Freq | 20 | 592.9 | 125.9 | 90 |
Low_Freq | 20 | 605.0 | 117.9 | 80 |
Pseudoword | 20 | 613.7 | 135.6 | 100 |
arrange() — Sort Rows Participant Condition RT_ms Accurate RT_s RT_log Fast_respons
1 6 Pseudoword 333.8950 TRUE 0.3338950 5.810826 TRUE
2 7 High_Freq 345.7743 TRUE 0.3457743 5.845786 TRUE
3 5 High_Freq 403.6127 TRUE 0.4036127 6.000456 TRUE
4 13 Pseudoword 441.0055 TRUE 0.4410055 6.089057 TRUE
5 1 Low_Freq 448.9416 TRUE 0.4489416 6.106893 TRUE Participant Condition RT_ms Accurate RT_s RT_log Fast_respons
1 18 Low_Freq 856.0582 TRUE 0.8560582 6.752338 FALSE
2 16 Pseudoword 845.5281 TRUE 0.8455281 6.739961 FALSE
3 15 High_Freq 790.6531 TRUE 0.7906531 6.672859 FALSE
4 19 Pseudoword 784.3431 TRUE 0.7843431 6.664847 FALSE
5 17 Low_Freq 782.4518 TRUE 0.7824518 6.662432 FALSErename() and relocate() Participant Condition ReactionTime Correct RT_s RT_log Fast_respons
1 1 High_Freq 555.4027 TRUE 0.5554027 6.319693 FALSE
2 1 Low_Freq 448.9416 TRUE 0.4489416 6.106893 TRUE
3 1 Pseudoword 499.9721 TRUE 0.4999721 6.214552 TRUEcount() — Quick Frequency Tables Condition n
1 High_Freq 20
2 Low_Freq 20
3 Pseudoword 20 Condition Accurate n
1 High_Freq FALSE 2
2 High_Freq TRUE 18
3 Low_Freq FALSE 4
4 Low_Freq TRUE 16
5 Pseudoword TRUE 20[1] 0 Participant Condition RT_ms Accurate RT_s RT_log
0 0 0 0 0 0
Fast_respons
0 Participant Condition RT_ms Accurate RT_s RT_log Fast_respons
1 1 High_Freq 555.4027 TRUE 0.5554027 6.319693 FALSE
2 1 Low_Freq 448.9416 TRUE 0.4489416 6.106893 TRUE
3 1 Pseudoword 499.9721 TRUE 0.4999721 6.214552 TRUE
4 2 High_Freq 514.8074 TRUE 0.5148074 6.243793 FALSE
5 2 Low_Freq 502.2348 TRUE 0.5022348 6.219068 FALSE
6 2 Pseudoword 474.1632 TRUE 0.4741632 6.161551 TRUE
7 3 High_Freq 563.1337 FALSE 0.5631337 6.333517 FALSE
8 3 Low_Freq 474.7938 FALSE 0.4747938 6.162881 TRUE
9 3 Pseudoword 591.0133 TRUE 0.5910133 6.381839 FALSE
10 4 High_Freq 476.5507 TRUE 0.4765507 6.166574 TRUE
11 4 Low_Freq 551.7678 FALSE 0.5517678 6.313127 FALSE
12 4 Pseudoword 605.7655 TRUE 0.6057655 6.406493 FALSE
13 5 High_Freq 403.6127 TRUE 0.4036127 6.000456 TRUE
14 5 Low_Freq 464.6666 FALSE 0.4646666 6.141320 TRUE
[ reached 'max' / getOption("max.print") -- omitted 46 rows ]Q1. What does dplyr::mutate() do?
Q2. You want the mean RT for each participant across all conditions. Which dplyr pipeline is correct?
What you’ll learn: How to create basic plots using ggplot2; the layered grammar of graphics
Key concept: Every ggplot2 plot is built by adding layers — data, aesthetics, geometries, and themes
ggplot2 is R’s most powerful and widely used plotting package. It is based on the Grammar of Graphics: the idea that every plot can be described by a consistent set of components.
Every ggplot2 plot has at least three components:
aes()): which variables map to which visual properties (x axis, y axis, colour, size, shape)geom_*()): how the data are visually represented (points, bars, lines, boxes)Additional optional components include scales, facets, themes, and labels.
ggplot(lex_data, aes(x = RT_ms, fill = Condition)) +
geom_histogram(bins = 20, color = "white", alpha = 0.7) +
facet_wrap(~ Condition, ncol = 1) +
scale_fill_manual(values = c("steelblue", "tomato", "seagreen")) +
theme_bw() +
theme(legend.position = "none", panel.grid.minor = element_blank()) +
labs(title = "Distribution of reaction times by condition",
x = "Reaction time (ms)", y = "Count") 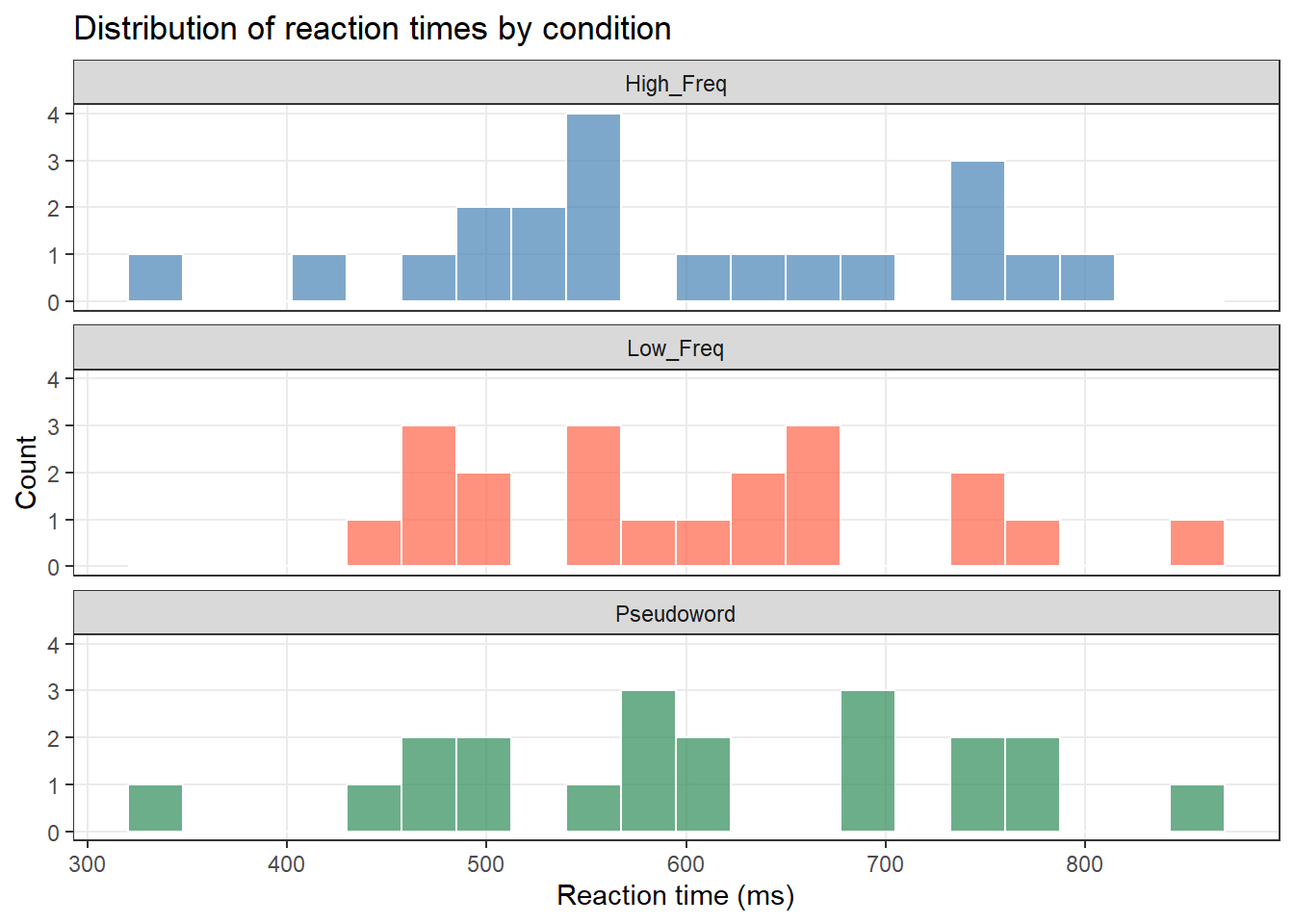
ggplot(lex_data, aes(x = Condition, y = RT_ms, fill = Condition)) +
geom_boxplot(alpha = 0.7, outlier.color = "gray40") +
stat_summary(fun = mean, geom = "point",
shape = 18, size = 3, color = "black") +
scale_fill_manual(values = c("steelblue", "tomato", "seagreen")) +
theme_bw() +
theme(legend.position = "none", panel.grid.minor = element_blank()) +
labs(title = "Reaction times by condition",
subtitle = "Diamond = group mean; box = median and IQR",
x = "Condition", y = "Reaction time (ms)") 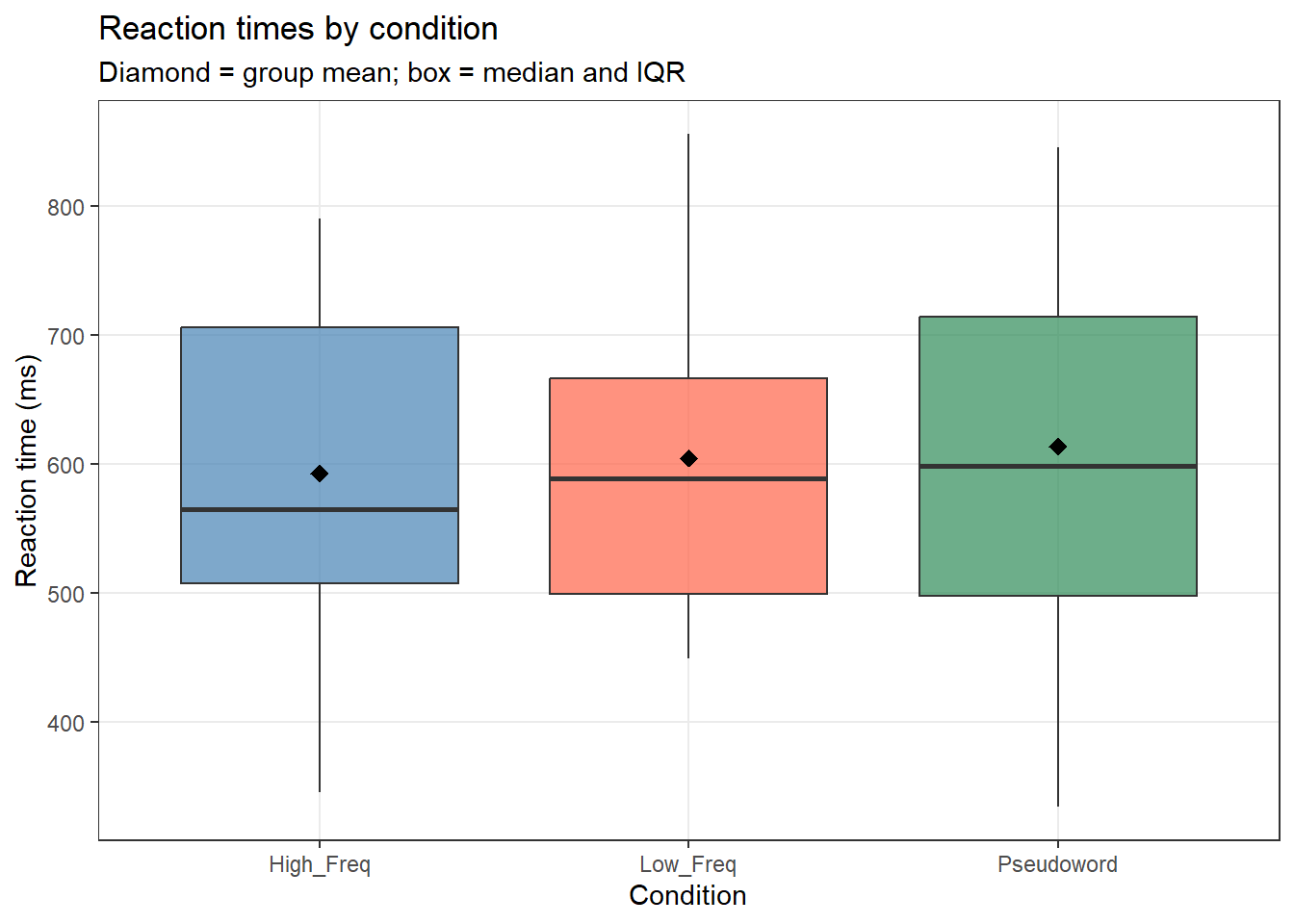
lex_data |>
dplyr::group_by(Condition) |>
dplyr::summarise(M_RT = mean(RT_ms),
SE = sd(RT_ms) / sqrt(n()),
.groups = "drop") |>
ggplot(aes(x = Condition, y = M_RT, fill = Condition)) +
geom_col(alpha = 0.8, width = 0.6) +
geom_errorbar(aes(ymin = M_RT - SE, ymax = M_RT + SE),
width = 0.2, linewidth = 0.8) +
scale_fill_manual(values = c("steelblue", "tomato", "seagreen")) +
theme_bw() +
theme(legend.position = "none", panel.grid.minor = element_blank()) +
labs(title = "Mean reaction time by condition",
subtitle = "Error bars = ±1 SE",
x = "Condition", y = "Mean RT (ms)") 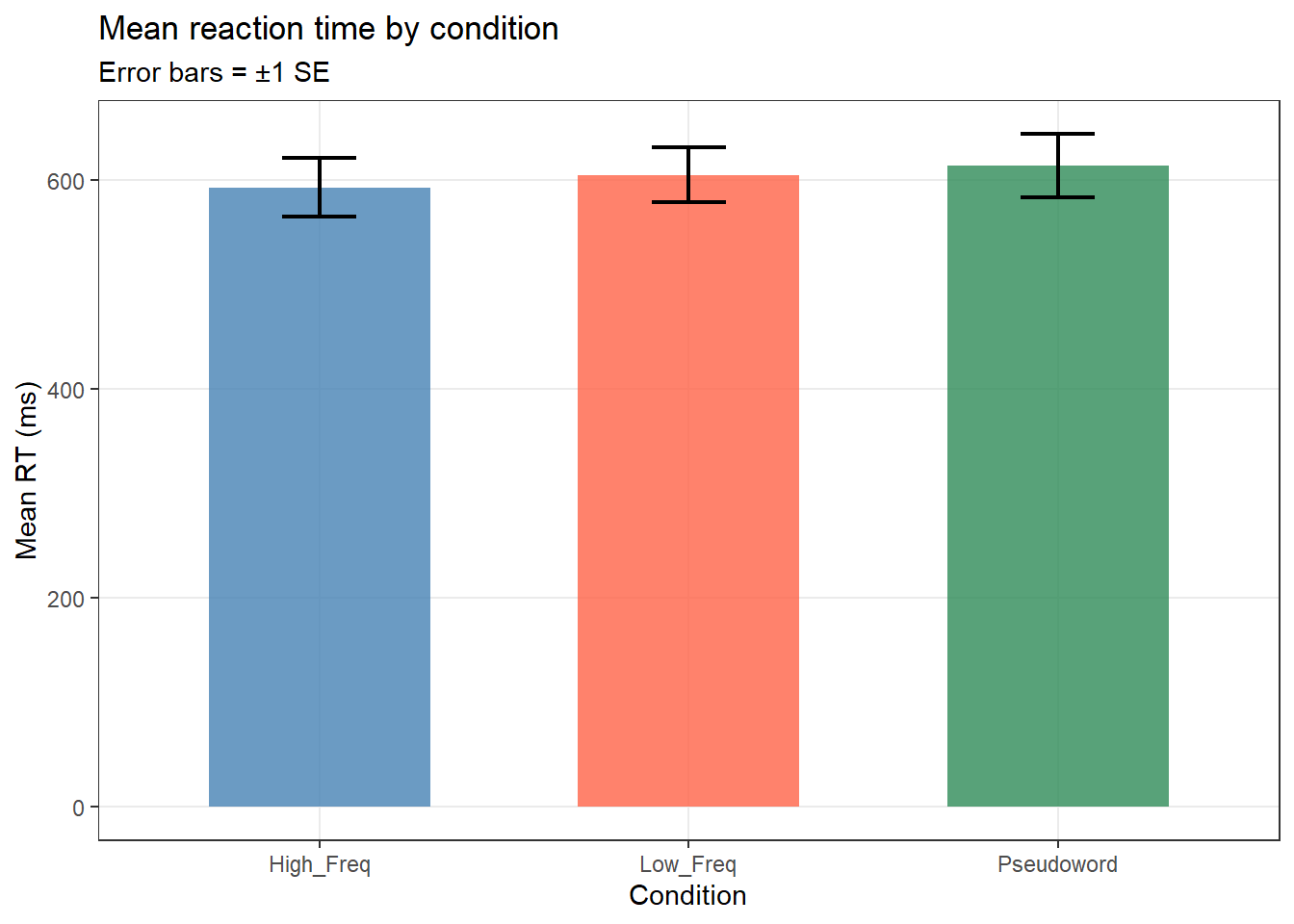
ggplot(lex_data, aes(x = Participant, y = RT_ms, color = Condition)) +
geom_point(alpha = 0.7, size = 2) +
scale_color_manual(values = c("steelblue", "tomato", "seagreen")) +
theme_bw() +
theme(panel.grid.minor = element_blank()) +
labs(title = "Individual RT observations by participant and condition",
x = "Participant ID", y = "Reaction time (ms)",
color = "Condition") 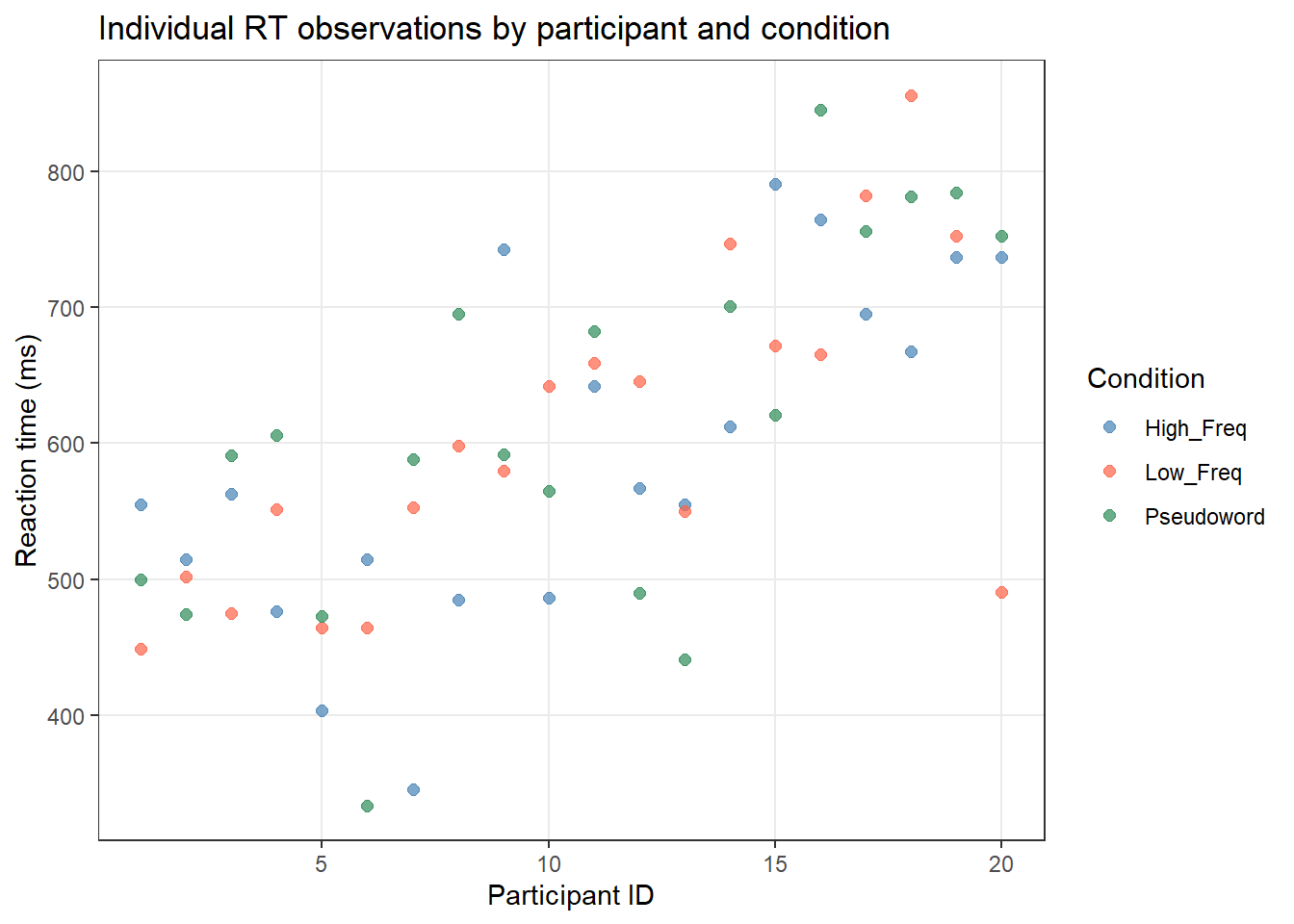
# Save the most recently displayed plot
ggsave(
filename = here::here("images", "my_plot.png"),
width = 8,
height = 5,
dpi = 300
)
# Save a named plot object
my_plot <- ggplot(lex_data, aes(x = RT_ms)) + geom_histogram()
ggsave(
plot = my_plot,
filename = here::here("images", "histogram.pdf"),
width = 6,
height = 4
) theme_bw() for a clean white background (LADAL standard)theme(panel.grid.minor = element_blank()) to remove minor gridlinesscale_color_manual() / scale_fill_manual() to control coloursfacet_wrap(~ variable) to create small multipleslabs() to set title, subtitle, and axis labels+ coord_flip() to swap x and y axes (useful for long category names)Q1. In ggplot2, what does aes() control?
Q2. Which geom_*() function would you use to create a histogram?
What you’ll learn: How to find help efficiently when you are stuck — both within R and online
Every R user gets stuck regularly. Knowing where to look for help is as important as knowing R itself.
RStudio’s Help tab (bottom right pane) renders help pages with formatted descriptions, argument lists, and examples.
Many packages include vignettes — detailed guides that show how to use the package end-to-end. These are often more useful than the function-level help pages:
Error messages are your friend — they tell you exactly what went wrong. Common error patterns:
object 'x' not found
→ The object x does not exist in your environment. Did you run the line that creates it? Is it spelled correctly (case-sensitive)?
could not find function "ggplot"
→ The package containing this function is not loaded. Did you run library(ggplot2)?
Error in read.csv("data.csv") : cannot open file
→ R cannot find the file. Check your working directory (getwd()), use here::here(), and check for typos in the filename.
non-numeric argument to binary operator
→ You tried to do arithmetic on a character string. Check the type of your object with class().
NAs introduced by coercion
→ R tried to convert a character to numeric but could not. The unconvertible values became NA. Inspect the affected column for unexpected text.
object of type 'closure' is not subsettable
→ You tried to index a function as if it were a data frame (e.g., mean[1]). Check whether you forgot parentheses somewhere.
The R community is enormous and helpful. When you encounter an error:
If you need to ask for help, always provide:
- A minimal reproducible example — the smallest piece of code that demonstrates the problem
- Your session info: sessionInfo()
- The exact error message (copy-paste, do not retype)
- What you expected to happen vs. what actually happened
The reprex package helps format reproducible examples: install.packages("reprex")
Resource | URL | Why useful |
|---|---|---|
R for Data Science | r4ds.hadley.nz | Free online book; the best comprehensive introduction to R and the tidyverse |
RStudio Cheatsheets | posit.co/resources/cheatsheets | One-page quick references for popular packages (dplyr, ggplot2, RMarkdown, etc.) |
CRAN Task Views | cran.r-project.org/web/views | Curated lists of R packages by topic (linguistics, NLP, spatial, etc.) |
Stack Overflow [r] | stackoverflow.com/questions/tagged/r | Answers to nearly every R question; search before posting |
Tidyverse documentation | tidyverse.org | Official documentation for dplyr, ggplot2, tidyr, readr, and more |
ggplot2 documentation | ggplot2.tidyverse.org | Function reference, articles, and extension gallery |
R Graph Gallery | r-graph-gallery.com | Hundreds of example plots with full reproducible code |
What you’ll learn: Habits and conventions that make your R code more readable, reproducible, and robust
Good coding habits matter more the longer your projects become. These practices are worth building from day one.
# This filters to English speakers onlyword_count not WordCount or wcx <- 5 * (3 + 2) not x<-5*(3+2)set.seed(42).Rproj)here::here() for all file paths — never hardcode absolute paths like "C:/Users/Martin/..."sessionInfo() to record package versionsrenv to snapshot your package environment# See all objects in your environment
ls()
# Remove a specific object
rm(my_temp_variable)
# Remove everything (use with caution!)
rm(list = ls())
# Check working directory
getwd()
# Change working directory (prefer R Projects over setwd())
setwd("path/to/folder") # avoid this; use R Projects instead Martin Schweinberger. 2026. Getting Started with R and RStudio. The Language Technology and Data Analysis Laboratory (LADAL), The University of Queensland, Australia. url: https://ladal.edu.au/tutorials/intror/intror.html (Version 2026.03.27), doi: 10.5281/zenodo.19242479.
@manual{martinschweinberger2026getting,
author = {Martin Schweinberger},
title = {Getting Started with R and RStudio},
year = {2026},
note = {https://ladal.edu.au/tutorials/intror/intror.html},
organization = {The Language Technology and Data Analysis Laboratory (LADAL), The University of Queensland, Australia},
edition = {2026.03.27}
doi = {10.5281/zenodo.19242479}
}R version 4.4.2 (2024-10-31 ucrt)
Platform: x86_64-w64-mingw32/x64
Running under: Windows 11 x64 (build 26200)
Matrix products: default
locale:
[1] LC_COLLATE=English_United States.utf8
[2] LC_CTYPE=English_United States.utf8
[3] LC_MONETARY=English_United States.utf8
[4] LC_NUMERIC=C
[5] LC_TIME=English_United States.utf8
time zone: Australia/Brisbane
tzcode source: internal
attached base packages:
[1] stats graphics grDevices datasets utils methods base
other attached packages:
[1] checkdown_0.0.13 flextable_0.9.7 here_1.0.1 tokenizers_0.3.0
[5] tm_0.7-16 NLP_0.3-2 readxl_1.4.3 quanteda_4.2.0
[9] tidytext_0.4.2 lubridate_1.9.4 forcats_1.0.0 stringr_1.5.1
[13] dplyr_1.2.0 purrr_1.0.4 readr_2.1.5 tidyr_1.3.2
[17] tibble_3.2.1 ggplot2_4.0.2 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] fastmatch_1.1-6 gtable_0.3.6 xfun_0.56
[4] htmlwidgets_1.6.4 lattice_0.22-6 tzdb_0.4.0
[7] vctrs_0.7.1 tools_4.4.2 generics_0.1.3
[10] parallel_4.4.2 janeaustenr_1.0.0 pkgconfig_2.0.3
[13] Matrix_1.7-2 data.table_1.17.0 RColorBrewer_1.1-3
[16] S7_0.2.1 uuid_1.2-1 lifecycle_1.0.5
[19] compiler_4.4.2 farver_2.1.2 textshaping_1.0.0
[22] codetools_0.2-20 litedown_0.9 fontLiberation_0.1.0
[25] fontquiver_0.2.1 SnowballC_0.7.1 htmltools_0.5.9
[28] yaml_2.3.10 pillar_1.10.1 openssl_2.3.2
[31] fontBitstreamVera_0.1.1 commonmark_2.0.0 stopwords_2.3
[34] zip_2.3.2 tidyselect_1.2.1 digest_0.6.39
[37] stringi_1.8.4 slam_0.1-55 labeling_0.4.3
[40] rprojroot_2.0.4 fastmap_1.2.0 grid_4.4.2
[43] cli_3.6.4 magrittr_2.0.3 withr_3.0.2
[46] gdtools_0.4.1 scales_1.4.0 timechange_0.3.0
[49] officer_0.6.7 rmarkdown_2.30 cellranger_1.1.0
[52] ragg_1.3.3 askpass_1.2.1 hms_1.1.3
[55] evaluate_1.0.3 knitr_1.51 markdown_2.0
[58] rlang_1.1.7 Rcpp_1.0.14 glue_1.8.0
[61] xml2_1.3.6 renv_1.1.1 rstudioapi_0.17.1
[64] jsonlite_1.9.0 R6_2.6.1 systemfonts_1.2.1 This tutorial was written with the assistance of Claude (claude.ai), a large language model created by Anthropic. Claude was used to draft and structure the entire tutorial, including all R code, conceptual explanations, and exercises. All content was reviewed and approved by Martin Schweinberger, who takes full responsibility for its accuracy.
---
title: "Getting Started with R and RStudio"
author: "Martin Schweinberger"
date: "2026"
params:
title: "Getting Started with R and RStudio"
author: "Martin Schweinberger"
year: "2026"
version: "2026.03.27"
url: "https://ladal.edu.au/tutorials/intror/intror.html"
institution: "The Language Technology and Data Analysis Laboratory (LADAL), The University of Queensland, Australia"
description: "This tutorial provides a hands-on introduction to R and RStudio for complete beginners, covering installation, the RStudio interface, basic R syntax, variables, functions, and writing a first R script. It is the recommended starting point for all LADAL users and serves as the prerequisite for all other practical tutorials in the collection."
doi: "10.5281/zenodo.19332886"
format:
html:
toc: true
toc-depth: 4
code-fold: show
code-tools: true
theme: cosmo
---
```{r setup, echo=FALSE, message=FALSE, warning=FALSE}
library(checkdown)
library(dplyr)
library(ggplot2)
library(tidyr)
library(flextable)
options(stringsAsFactors = FALSE)
options(scipen = 100)
options(max.print = 100)
```
{width="100%" height="200px" loading="lazy"}
# Introduction {#intro}
{ width=15% style="float:right; padding:10px" }
This tutorial introduces **R and RStudio** — the programming language and development environment used throughout LADAL. It is aimed at complete beginners with no prior programming experience, and walks through everything you need to get up and running: installing software, understanding the RStudio interface, setting up a reproducible project, and working with R for the first time.
R is a free, open-source programming language designed specifically for data analysis and statistics. It is the most widely used tool for quantitative research in linguistics, the social sciences, and the digital humanities — and for good reason. R gives you complete control over your analysis, produces publication-quality graphics, and keeps your work fully transparent and reproducible.
This tutorial will not turn you into an expert. Its goal is to give you a solid, well-structured foundation: to know *where things are*, *how to think about R*, and *how to start doing real things with data*. The rest of LADAL's tutorials build from here.
::: {.callout-note}
## Prerequisite Tutorials
This tutorial has no prerequisites — it is designed for complete beginners. However, the following background tutorials are helpful companions:
- [Introduction to Quantitative Reasoning](/tutorials/introquant/introquant.html)
- [Basic Concepts in Quantitative Research](/tutorials/basicquant/basicquant.html)
- [Reproducible Research](/tutorials/repro/repro.html)
:::
::: {.callout-tip}
## What This Tutorial Covers
1. **Installing R and RStudio** — getting everything set up on your computer
2. **The RStudio interface** — understanding the four panes and how to navigate them
3. **R Projects and R Notebooks** — setting up reproducible, well-organised workflows
4. **R fundamentals** — objects, functions, operators, and data types
5. **Data structures** — vectors, data frames, lists, and factors
6. **Indexing and subsetting** — accessing and filtering data
7. **Working with data** — loading, inspecting, and manipulating tabular data
8. **Basic visualisation** — creating your first plots with `ggplot2`
9. **Getting help** — where to turn when things go wrong
:::
::: {.callout-note}
## Citation
```{r citation-callout-top, echo=FALSE, results='asis'}
cat(
params$author, ". ",
params$year, ". *",
params$title, "*. ",
params$institution, ". ",
"url: ", params$url, " ",
"(Version ", params$version, "), ",
"doi: ", params$doi, ".",
sep = ""
)
```
:::
---
## Why R? {-}
Before diving in, it is worth briefly explaining why R is worth learning.
R is **free and open-source** — there are no licensing costs, ever. It is the **dominant tool** for statistical analysis in linguistics, psychology, and the social sciences. It has a vast ecosystem of over 20,000 contributed packages that extend its capabilities to cover almost any analytical task imaginable. Its **reproducibility** features — the ability to combine code, output, and prose in a single document — mean your analyses can be fully transparent and re-run by anyone. And its **visualisation** capabilities, particularly through `ggplot2`, are unmatched.
The learning curve is real but manageable. This tutorial gives you the foundation you need.
---
## Preparation and Session Set-up {-}
Install the packages used in this tutorial (only needed once):
```{r install, echo=T, eval=F, message=FALSE, warning=FALSE}
install.packages("dplyr")
install.packages("ggplot2")
install.packages("tidyr")
install.packages("flextable")
install.packages("readxl")
install.packages("here")
install.packages("checkdown")
```
Load the packages at the start of each session:
```{r load, echo=T, eval=T, message=FALSE, warning=FALSE}
library(dplyr) # data manipulation
library(ggplot2) # data visualisation
library(tidyr) # data reshaping
library(flextable) # formatted tables
library(here) # robust file paths
library(checkdown) # interactive exercises
```
---
# Installing R and RStudio {#install}
::: {.callout-note}
## Section Overview
**What you'll learn:** How to install R and RStudio on your computer
**Why it matters:** You need both installed to follow any LADAL tutorial
**Time:** ~15–30 minutes (mostly waiting for downloads)
:::
R and RStudio are two separate pieces of software that work together. Think of **R** as the engine and **RStudio** as the car — you need both, and you interact almost exclusively with RStudio.
## Installing R {-}
R must be installed before RStudio. Visit [**cran.r-project.org**](https://cran.r-project.org/) and select the download for your operating system:
- **Windows**: click *Download R for Windows* → *base* → *Download R x.x.x for Windows*
- **Mac**: click *Download R for macOS* → select the version matching your macOS
- **Linux**: follow the instructions for your distribution
Run the downloaded installer and accept the default settings throughout.
::: {.callout-tip}
## Keeping R Up to Date
R releases a new version approximately once a year. To check your current version, run `R.version$version.string` in the console. To update on Windows, the `installr` package automates the process:
```{r update_r, eval=FALSE}
install.packages("installr")
library(installr)
updateR()
```
On Mac, download the new version from CRAN and install over the existing version.
:::
## Installing RStudio {-}
Visit [**posit.co/download/rstudio-desktop**](https://posit.co/download/rstudio-desktop/) and download the free **RStudio Desktop** version for your operating system. Run the installer and accept the defaults.
After installation, open **RStudio** (not R directly). RStudio will automatically detect your R installation.
---
# The RStudio Interface {#interface}
::: {.callout-note}
## Section Overview
**What you'll learn:** How to navigate the four panes of RStudio and what each one does
**Key concept:** The difference between the Console (run immediately) and the Script Editor (save and reuse)
:::
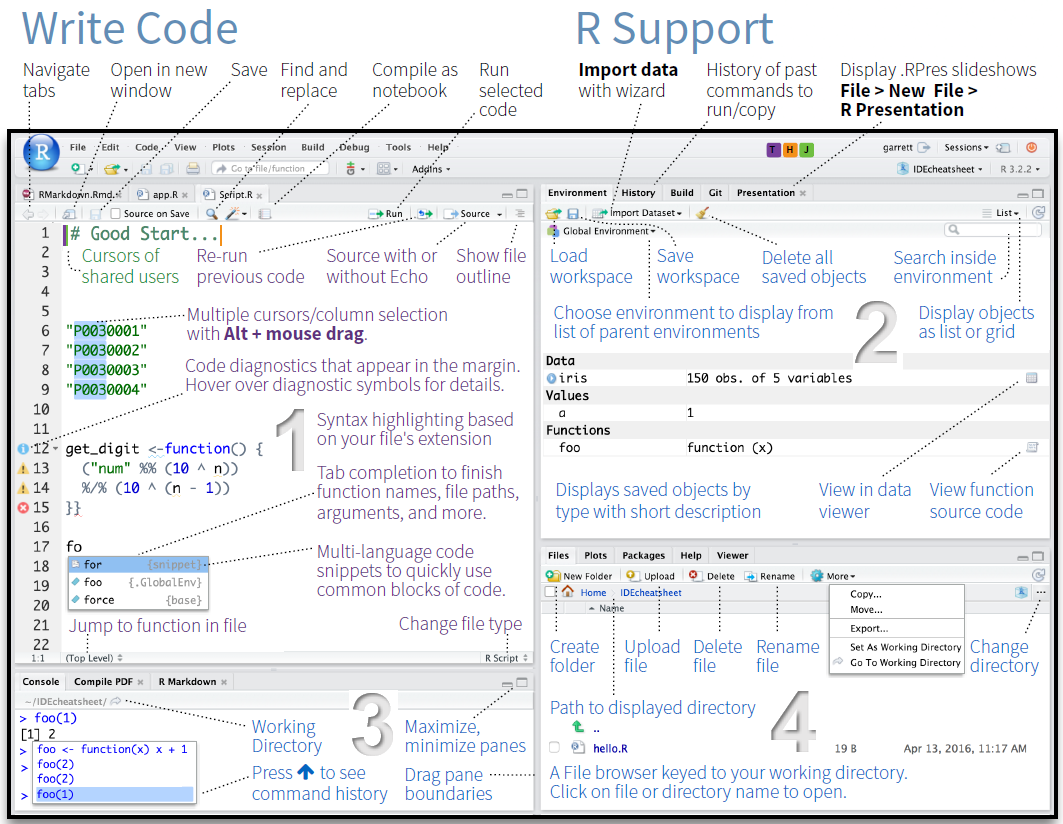
When you first open RStudio, you will see an interface divided into panes. The screenshot below shows a typical RStudio session with all four panes visible.
{ width=100% }
RStudio has four main panes:
## Pane 1: Script Editor (top left) {-}
This is where you **write and save code**. Code typed here does not run automatically — you must explicitly execute it. This is where all your analysis lives.
To run a line of code from the Script Editor, place your cursor on that line and press `Ctrl + Enter` (Windows/Linux) or `Cmd + Enter` (Mac). To run a highlighted block, select the code first and then press the same shortcut.
## Pane 2: Console (bottom left) {-}
This is where R **executes code and displays text output**. When you run code from the Script Editor, it appears here. You can also type directly into the Console and press `Enter` to run commands immediately.
Use the Console for quick experiments. Use the Script Editor for anything you want to keep.
::: {.callout-tip}
## Console Shortcuts
- Press the **Up arrow** in the Console to recall previous commands
- Type the beginning of a command and press `Tab` to autocomplete
- Type `?function_name` to open the help page for any function
:::
## Pane 3: Environment and History (top right) {-}
The **Environment** tab shows all objects currently loaded in your R session — data frames, variables, vectors, and so on. Clicking on a data frame here opens a spreadsheet-style viewer.
The **History** tab logs all commands you have run in the current session.
## Pane 4: Files, Plots, Help, Packages (bottom right) {-}
This multi-tab pane contains:
- **Files**: Browse your project folder
- **Plots**: View graphics output here
- **Help**: Documentation for functions and packages (also accessible via `?`)
- **Packages**: See which packages are installed and loaded
- **Viewer**: Preview rendered documents
---
# Projects and Notebooks {#projects}
::: {.callout-note}
## Section Overview
**What you'll learn:** How to set up a reproducible project in RStudio; what an R Notebook is and why to use one
**Key concept:** An R Project keeps all your files, code, and data together in one self-contained folder
:::
Good organisation before you start coding saves a great deal of trouble later. This section walks through the recommended setup.
## Step 1: Create a Project Folder {-}
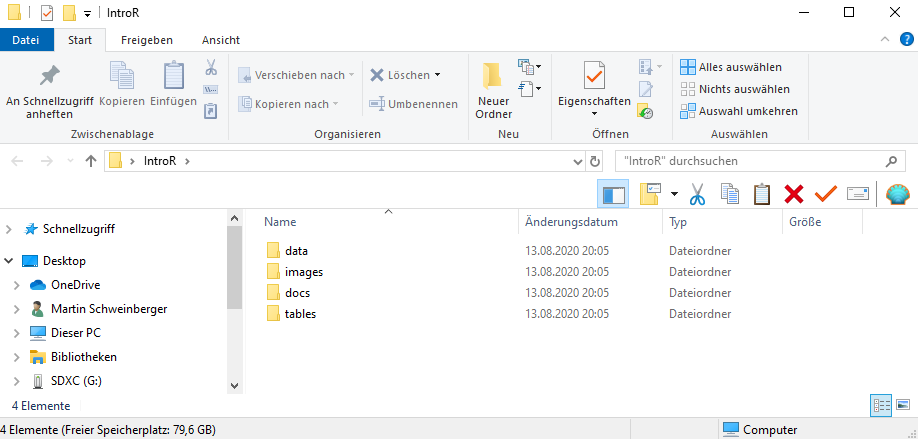
Before opening RStudio, create a folder on your computer for your project. Inside it, create the following sub-folders:
```
my_project/
├── data/ ← raw and processed data files
├── images/ ← figures saved from R
├── tables/ ← tables exported from R
└── docs/ ← notes, reports, and output documents
```
{ width=75% }
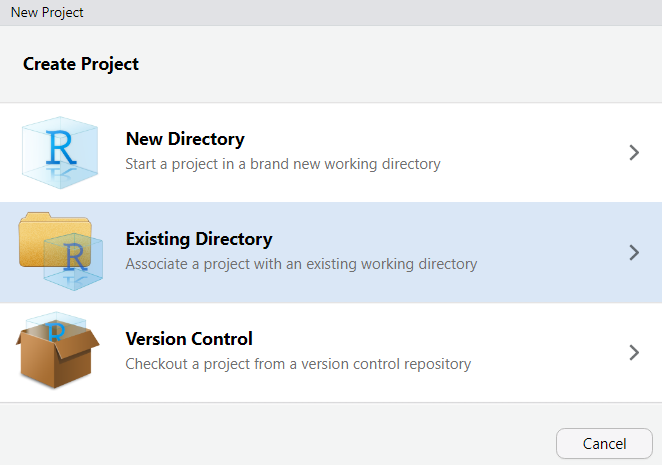
## Step 2: Create an R Project {-}
An **R Project** tells RStudio that a folder is a self-contained project. It sets the **working directory** automatically (so file paths are predictable) and keeps your project's history and settings separate from other projects.
To create an R Project:
1. Open RStudio
2. Click `File` → `New Project`
3. Select `Existing Directory`
4. Navigate to your project folder and click `Create Project`
RStudio will restart and you will see your project name in the top-right corner. You are now working inside your project.
{ width=35% }
::: {.callout-important}
## Always Work Inside an R Project
When you open RStudio, always open your project first (either by double-clicking the `.Rproj` file in your folder, or via `File → Open Project`). This ensures file paths work correctly and your environment is isolated.
:::
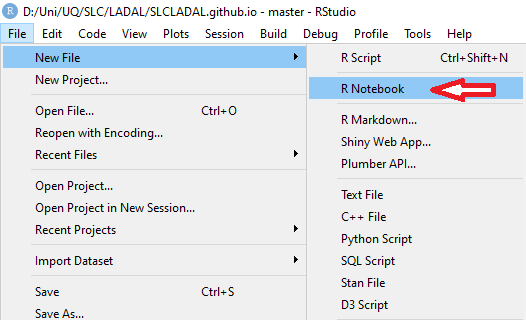
## Step 3: Create an R Notebook {-}
An **R Notebook** (`.Rmd` or `.qmd` file) combines prose, code, and output in a single document. This is the standard format for LADAL tutorials and is highly recommended for your own analyses — it keeps your thinking and your code together.
To create an R Notebook:
1. Click `File` → `New File` → `R Notebook`
2. Give it a meaningful title
3. Save it in your project folder
{ width=50% }
The notebook uses **R Markdown** — a simple formatting syntax explained below.
## R Markdown Basics {-}
R Markdown lets you write formatted prose alongside executable code. Here is a quick reference:
```
# Heading 1
## Heading 2
### Heading 3
**bold text**
*italic text*
`inline code`
- bullet point
- another bullet
1. numbered item
2. another item
[link text](https://url.com)
```
Code is written inside **code chunks** (fenced with triple backticks):
````
```{r chunk-name, message=FALSE, warning=FALSE}
# your R code here
2 + 2
```
````
When you click **Knit** (or **Render** in Quarto), R Markdown executes all code chunks and weaves the output together with your prose into a finished HTML, PDF, or Word document.
::: {.callout-tip}
## Reproducibility
The power of R Notebooks is reproducibility: your entire analysis — every number, table, and figure — is regenerated from scratch each time you render the document. Anyone with your `.Rmd` file and data can reproduce your results exactly.
:::
---
# R Fundamentals {#fundamentals}
::: {.callout-note}
## Section Overview
**What you'll learn:** The core building blocks of R — objects, functions, operators, and assignment
**Key concepts:** Everything in R is an object; everything you do in R uses a function
:::
## Setting Up a Session {-}
At the top of any script or notebook, set global options and load packages. This makes your session reproducible from the very first line.
```{r session_setup, message=FALSE, warning=FALSE}
# Global options
options(stringsAsFactors = FALSE) # keep character variables as text
options(scipen = 100) # avoid scientific notation
options(max.print = 100) # limit printed output
# Load packages
library(dplyr)
library(ggplot2)
```
## Objects and Assignment {-}
In R, everything is stored as an **object**. You create objects using the **assignment operator** `<-`:
```{r objects}
# Create a numeric object
my_number <- 42
# Create a character (text) object
my_name <- "linguistics"
# Create a logical object
is_true <- TRUE
# View an object by typing its name
my_number
my_name
is_true
```
::: {.callout-tip}
## Naming Objects
Good object names are:
- **lowercase** with underscores for spaces: `word_count`, not `Word Count`
- **descriptive**: `reaction_time_ms` is better than `x`
- **not starting with a number**: `data1` is valid; `1data` is not
- **not reserved words**: don't use `c`, `t`, `df`, `mean`, `TRUE`, `FALSE`, `NULL` as object names
R is **case-sensitive**: `MyData` and `mydata` are different objects.
:::
## Functions {-}
A **function** takes one or more inputs (called **arguments**), does something, and returns an output. Functions are called by name followed by parentheses containing the arguments:
```{r functions}
# sqrt() takes a number and returns its square root
sqrt(144)
# round() rounds a number to a specified number of decimal places
round(3.14159, digits = 2)
# nchar() counts the characters in a string
nchar("linguistics")
# paste() joins strings together
paste("language", "data", "analysis", sep = "-")
```
You can nest functions — the inner function runs first:
```{r nested_functions}
# Round the square root of 2 to 3 decimal places
round(sqrt(2), digits = 3)
```
## Operators {-}
R provides standard arithmetic and logical operators:
```{r operators}
# Arithmetic operators
10 + 3 # addition
10 - 3 # subtraction
10 * 3 # multiplication
10 / 3 # division
10 ^ 2 # exponentiation
10 %% 3 # modulo (remainder)
```
```{r logical_ops}
# Comparison operators (return TRUE or FALSE)
5 > 3 # greater than
5 < 3 # less than
5 == 5 # equal to (note: double equals!)
5 != 3 # not equal to
5 >= 5 # greater than or equal to
# Logical operators
TRUE & FALSE # AND
TRUE | FALSE # OR
!TRUE # NOT
```
::: {.callout-warning}
## `=` vs `==`
One of the most common beginner errors: `=` is used for assignment (interchangeable with `<-` in most cases, though `<-` is preferred); `==` tests whether two things are equal. `5 = 3` will produce an error; `5 == 3` returns `FALSE`.
:::
---
::: {.callout-tip}
## Exercises: R Fundamentals
:::
**Q1. What does the assignment operator `<-` do?**
```{r}
#| echo: false
#| label: "FUND_Q1"
check_question("It creates an object by storing a value under a name in the current environment",
options = c(
"It creates an object by storing a value under a name in the current environment",
"It tests whether two values are equal",
"It subtracts the right-hand value from the left-hand value",
"It calls a function with the specified argument"
),
type = "radio",
q_id = "FUND_Q1",
random_answer_order = TRUE,
button_label = "Check answer",
right = "Correct! x <- 42 creates an object named x and stores the value 42 in it. From that point on, typing x anywhere in your code returns 42. The shortcut for <- in RStudio is Alt + - (Windows) or Option + - (Mac).",
wrong = "Think about what happens after you write x <- 42 and then type x — what does R show you?")
```
---
**Q2. You run `my_var <- 10`. What will `my_var * 3 + 1` return?**
```{r}
#| echo: false
#| label: "FUND_Q2"
check_question("31",
options = c("31", "30", "13", "An error, because my_var is not a function"),
type = "radio",
q_id = "FUND_Q2",
random_answer_order = FALSE,
button_label = "Check answer",
right = "Correct! R substitutes the stored value: 10 * 3 + 1 = 30 + 1 = 31. Standard mathematical order of operations applies.",
wrong = "Remember that my_var holds the value 10. R replaces my_var with 10 and then evaluates: 10 * 3 + 1.")
```
---
**Q3. Which of the following is NOT a valid object name in R?**
```{r}
#| echo: false
#| label: "FUND_Q3"
check_question("2nd_group",
options = c("2nd_group", "group_2", "group.two", "myGroup"),
type = "radio",
q_id = "FUND_Q3",
random_answer_order = FALSE,
button_label = "Check answer",
right = "Correct! Object names in R cannot start with a digit. 2nd_group would throw a syntax error. group_2, group.two (dots are allowed), and myGroup are all valid — though the LADAL style convention is lowercase with underscores (group_2).",
wrong = "Which option begins with something other than a letter or dot?")
```
---
# Data Types {#datatypes}
::: {.callout-note}
## Section Overview
**What you'll learn:** The six basic data types in R and why they matter
**Key concept:** The type of your data determines which operations are valid
:::
Every object in R has a **type** (also called a **class**). The four types you will encounter most often are:
```{r datatypes}
# Numeric (continuous numbers)
age <- 28.5
class(age)
# Integer (whole numbers; the L suffix forces integer type)
count <- 42L
class(count)
# Character (text; always in quotes)
language <- "English"
class(language)
# Logical (TRUE or FALSE only)
is_native <- TRUE
class(is_native)
```
You can check the type of any object with `class()` or `typeof()`, and test for specific types:
```{r type_tests}
is.numeric(age)
is.character(language)
is.logical(is_native)
```
You can **convert** between types using coercion functions:
```{r coercion}
# Character to numeric
as.numeric("3.14")
# Numeric to character
as.character(42)
# Numeric to logical (0 = FALSE, everything else = TRUE)
as.logical(0)
as.logical(1)
as.logical(-99)
```
::: {.callout-warning}
## Coercion Failures
When R cannot coerce a value, it introduces `NA` (missing value) with a warning:
```{r coerce_fail, warning=TRUE}
as.numeric("hello") # "hello" cannot be a number → NA
```
`NA` stands for *Not Available* and represents missing data. It propagates through calculations — any arithmetic involving `NA` returns `NA` unless specifically handled.
:::
---
# Data Structures {#structures}
::: {.callout-note}
## Section Overview
**What you'll learn:** How R organises collections of data — vectors, data frames, lists, and factors
**Key concept:** Vectors are the fundamental unit; data frames are collections of equal-length vectors
:::
## Vectors {-}
A **vector** is a sequence of values of the *same type*. Vectors are created with `c()` (short for *combine*):
```{r vectors}
# Numeric vector
word_lengths <- c(3, 5, 2, 8, 4, 6, 1)
# Character vector
languages <- c("English", "German", "Mandarin", "Arabic")
# Logical vector
is_content_word <- c(TRUE, TRUE, FALSE, TRUE, FALSE)
```
You can perform operations on entire vectors at once — R applies them element-by-element:
```{r vector_ops}
# Arithmetic on a vector
word_lengths * 2
# Logical comparison on a vector
word_lengths > 4
# Common summary functions
length(word_lengths) # number of elements
sum(word_lengths) # sum
mean(word_lengths) # mean
sd(word_lengths) # standard deviation
min(word_lengths) # minimum
max(word_lengths) # maximum
range(word_lengths) # min and max together
```
### Sequences and Repetitions {-}
```{r sequences}
# Create a sequence with :
1:10
# Create a sequence with seq()
seq(from = 0, to = 1, by = 0.25)
seq(from = 1, to = 100, length.out = 5)
# Repeat values with rep()
rep("yes", times = 3)
rep(c("A", "B"), times = 4)
rep(c("A", "B"), each = 4)
```
## Factors {-}
A **factor** is a special type of vector for **categorical variables**. Factors have a fixed set of levels (categories) and are essential for grouping in analyses and plots.
```{r factors}
# Create a factor
register <- factor(c("Formal", "Informal", "Formal", "ReadAloud", "Informal"))
# Inspect the factor
register
levels(register) # the unique categories
nlevels(register) # how many categories
table(register) # frequency of each level
```
By default, levels are ordered alphabetically. You can specify a custom order:
```{r factor_levels}
# Custom level order (important for plots and models)
register_ordered <- factor(
c("Formal", "Informal", "Formal", "ReadAloud", "Informal"),
levels = c("Formal", "ReadAloud", "Informal")
)
levels(register_ordered)
```
## Data Frames {-}
A **data frame** is R's equivalent of a spreadsheet — a table where each column is a vector of the same length. Data frames are the most common way to store linguistic data.
```{r dataframes}
# Create a data frame from scratch
speakers <- data.frame(
ID = 1:6,
Name = c("Alice", "Bob", "Carol", "David", "Eve", "Frank"),
L1 = c("English", "German", "English", "Mandarin", "English", "Arabic"),
Age = c(24, 31, 28, 22, 35, 27),
Proficiency = factor(c("Advanced", "Intermediate", "Advanced",
"Beginner", "Intermediate", "Advanced"),
levels = c("Beginner", "Intermediate", "Advanced"))
)
# Inspect the data frame
speakers
```
Key functions for inspecting a data frame:
```{r df_inspect}
nrow(speakers) # number of rows (observations)
ncol(speakers) # number of columns (variables)
dim(speakers) # both at once
names(speakers) # column names
str(speakers) # structure: types and first values
head(speakers, n = 3) # first 3 rows
tail(speakers, n = 2) # last 2 rows
summary(speakers) # summary statistics per column
```
## Lists {-}
A **list** is the most flexible data structure — it can hold objects of *different types and lengths*, including other lists.
```{r lists}
# Create a list with mixed types
my_list <- list(
name = "Study 1",
n = 30,
groups = c("Control", "Treatment"),
complete = TRUE
)
# Access list elements with $ or [[]]
my_list$name
my_list[["n"]]
```
Lists are commonly returned by statistical model functions (e.g., `lm()` returns a list). You rarely create them from scratch but frequently need to extract elements from them.
---
::: {.callout-tip}
## Exercises: Data Structures
:::
**Q1. You run `x <- c(1, 2, "three", 4)`. What type will x be?**
```{r}
#| echo: false
#| label: "STR_Q1"
check_question("Character — R coerces all elements to the most flexible type that can represent all values",
options = c(
"Character — R coerces all elements to the most flexible type that can represent all values",
"Numeric — the numbers override the character",
"Mixed — R keeps each element as its original type",
"It produces an error because you cannot mix types in a vector"
),
type = "radio",
q_id = "STR_Q1",
random_answer_order = TRUE,
button_label = "Check answer",
right = 'Correct! Vectors must contain one type only. When you mix types, R silently coerces everything to the most general type that can represent all values. The hierarchy is: logical → integer → numeric → character. Because "three" cannot be numeric, everything is coerced to character: c("1", "2", "three", "4"). This is called implicit coercion and is a common source of surprising results.',
wrong = 'Vectors in R are homogeneous — they can only hold one type. What happens when you try to put "three" into a numeric vector?')
```
---
**Q2. What is the difference between a factor and a character vector?**
```{r}
#| echo: false
#| label: "STR_Q2"
check_question("A factor has a fixed set of predefined levels (categories); a character vector is just text with no inherent structure",
options = c(
"A factor has a fixed set of predefined levels (categories); a character vector is just text with no inherent structure",
"Factors can only contain numbers; character vectors contain text",
"There is no practical difference — they behave identically",
"Character vectors are faster to compute with than factors"
),
type = "radio",
q_id = "STR_Q2",
random_answer_order = TRUE,
button_label = "Check answer",
right = "Correct! A factor stores categorical data as integers internally, with a levels attribute recording what each integer represents. This means factors have a defined set of valid categories, can be ordered, and are handled correctly in statistical models and plots (e.g., as a grouping variable). A plain character vector has no such structure — R treats each unique string independently with no notion of grouping.",
wrong = "Think about what makes categorical data special in statistics. What does it mean for a variable to have predefined categories?")
```
---
**Q3. What does `dim(df)` return for a data frame with 50 rows and 4 columns?**
```{r}
#| echo: false
#| label: "STR_Q3"
check_question("c(50, 4) — a vector with number of rows first, then number of columns",
options = c(
"c(50, 4) — a vector with number of rows first, then number of columns",
"c(4, 50) — columns first, then rows",
"200 — the total number of cells",
"A list with named elements $rows and $cols"
),
type = "radio",
q_id = "STR_Q3",
random_answer_order = TRUE,
button_label = "Check answer",
right = "Correct! dim() always returns a two-element vector in the order (rows, columns). This convention — rows before columns — is consistent throughout R: matrix notation, subsetting, and model output all follow [row, column] order. nrow() and ncol() give each dimension separately.",
wrong = "R consistently uses rows-before-columns ordering. Check: what does dim() return, and in what order?")
```
---
# Indexing and Subsetting {#indexing}
::: {.callout-note}
## Section Overview
**What you'll learn:** How to access specific elements, rows, columns, and subsets of your data
**Key concept:** Square brackets `[ ]` select by position; `$` selects columns by name; `dplyr` verbs filter by condition
:::
Extracting exactly the data you need is one of the most fundamental R skills.
## Indexing Vectors {-}
Use square brackets `[ ]` with a position number (index) to extract elements from a vector. **R indexing starts at 1** (not 0 as in Python).
```{r vector_index}
languages <- c("English", "German", "Mandarin", "Arabic", "French")
# Extract a single element
languages[1] # first element
languages[4] # fourth element
# Extract multiple elements
languages[c(1, 3)] # first and third
languages[2:4] # second through fourth
# Exclude elements (negative indexing)
languages[-2] # everything except the second element
languages[-c(1,5)] # everything except first and fifth
# Logical indexing
word_lengths <- c(3, 5, 2, 8, 4, 6, 1)
word_lengths[word_lengths > 4] # elements greater than 4
word_lengths[word_lengths == min(word_lengths)] # the minimum value
```
## Indexing Data Frames {-}
Data frames have two dimensions: `df[row, column]`. Leave one blank to select all rows or all columns.
```{r df_index}
# Using the speakers data frame from earlier
# Single cell: row 2, column 3
speakers[2, 3]
# Entire row 1
speakers[1, ]
# Entire column 3 (returns a vector)
speakers[, 3]
# Column by name using $
speakers$Age
speakers$L1
# Multiple rows and columns
speakers[1:3, c("Name", "Age")]
```
## Subsetting with `dplyr` {-}
While base R indexing works, the `dplyr` package provides **cleaner, more readable** syntax for filtering and selecting data. These are the two most important `dplyr` verbs for subsetting:
```{r dplyr_subset}
# filter() keeps rows that meet a condition
speakers |>
dplyr::filter(L1 == "English")
# select() keeps specified columns
speakers |>
dplyr::select(Name, Age, Proficiency)
# Combine both
speakers |>
dplyr::filter(Age < 30) |>
dplyr::select(Name, L1, Age)
```
::: {.callout-tip}
## The Pipe Operator `|>`
The pipe `|>` (from the `magrittr`/`dplyr` packages) passes the result on the left to the function on the right. It lets you chain operations in a readable left-to-right sequence instead of nesting functions:
```r
# Without pipe (hard to read)
select(filter(speakers, Age < 30), Name, Age)
# With pipe (reads like a sentence)
speakers |> filter(Age < 30) |> select(Name, Age)
```
R 4.1+ also has a native pipe `|>` that works similarly. LADAL tutorials use `|>`.
:::
---
::: {.callout-tip}
## Exercises: Indexing
:::
**Q1. Given `v <- c(10, 20, 30, 40, 50)`, what does `v[c(2, 4)]` return?**
```{r}
#| echo: false
#| label: "IDX_Q1"
check_question("20 40",
options = c("20 40", "10 30 50", "2 4", "An error"),
type = "radio",
q_id = "IDX_Q1",
random_answer_order = FALSE,
button_label = "Check answer",
right = "Correct! c(2, 4) is an index vector selecting the 2nd and 4th elements of v. v[2] = 20 and v[4] = 40, so v[c(2, 4)] returns c(20, 40).",
wrong = "Remember: the numbers inside the square brackets are *positions*, not values. c(2, 4) means 'give me the element at position 2 and the element at position 4'.")
```
---
**Q2. How do you use `dplyr::filter()` to keep only rows where the column `Proficiency` equals `"Advanced"`?**
```{r}
#| echo: false
#| label: "IDX_Q2"
check_question('df |> dplyr::filter(Proficiency == "Advanced")',
options = c(
'df |> dplyr::filter(Proficiency == "Advanced")',
'df |> dplyr::filter(Proficiency = "Advanced")',
'df |> dplyr::select(Proficiency == "Advanced")',
'df[df$Proficiency = "Advanced", ]'
),
type = "radio",
q_id = "IDX_Q2",
random_answer_order = TRUE,
button_label = "Check answer",
right = 'Correct! filter() keeps rows where the condition is TRUE. The condition uses == (double equals) for equality testing — a single = would be a syntax error inside filter(). select() picks columns, not rows, so that option is wrong. The base R indexing option uses = instead of ==, which would also error.',
wrong = 'There are two things to check: which function filters rows vs. columns, and which operator tests equality vs. assignment.')
```
---
# Working with Data {#data}
::: {.callout-note}
## Section Overview
**What you'll learn:** How to load data from files, inspect it, and perform common data manipulation operations
**Key functions:** `read.csv()`, `readxl::read_excel()`, `dplyr::mutate()`, `dplyr::group_by()`, `dplyr::summarise()`
:::
## Loading Data {-}
### From CSV {-}
```{r load_csv, eval=FALSE}
# Base R
my_data <- read.csv("data/my_file.csv")
# Using here() for robust paths (recommended)
my_data <- read.csv(here::here("data", "my_file.csv"))
# Tidyverse readr (slightly faster, better defaults)
my_data <- readr::read_csv(here::here("data", "my_file.csv"))
```
### From Excel {-}
```{r load_excel, eval=FALSE}
library(readxl)
my_data <- readxl::read_excel(here::here("data", "my_file.xlsx"))
# Specify a sheet
my_data <- readxl::read_excel(here::here("data", "my_file.xlsx"), sheet = "Sheet2")
```
### Saving Data {-}
```{r save_data, eval=FALSE}
# Save as CSV
write.csv(my_data, here::here("data", "processed_data.csv"), row.names = FALSE)
# Save as R object (preserves factors and other R-specific attributes)
saveRDS(my_data, here::here("data", "processed_data.rds"))
# Load an RDS file
my_data <- readRDS(here::here("data", "processed_data.rds"))
```
## Manipulating Data with dplyr {-}
We will use a simulated linguistic dataset to demonstrate the key `dplyr` operations. The dataset contains reaction times and accuracy from a lexical decision task:
```{r create_data}
set.seed(42)
n <- 60
lex_data <- data.frame(
Participant = rep(1:20, each = 3),
Condition = rep(c("High_Freq", "Low_Freq", "Pseudoword"), times = 20),
RT_ms = c(
rnorm(20, mean = 480, sd = 55), # High frequency: fast
rnorm(20, mean = 610, sd = 70), # Low frequency: slower
rnorm(20, mean = 730, sd = 80) # Pseudowords: slowest
),
Accurate = sample(c(TRUE, FALSE), n, replace = TRUE, prob = c(0.9, 0.1))
) |>
dplyr::mutate(Condition = factor(Condition,
levels = c("High_Freq", "Low_Freq", "Pseudoword")))
```
### `mutate()` — Add or Modify Columns {-}
```{r mutate}
# Add a new column converting RT to seconds
lex_data <- lex_data |>
dplyr::mutate(
RT_s = RT_ms / 1000,
RT_log = log(RT_ms),
Fast_respons = RT_ms < 500
)
head(lex_data)
```
### `group_by()` and `summarise()` — Aggregate by Group {-}
```{r summarise}
lex_data |>
dplyr::group_by(Condition) |>
dplyr::summarise(
n = n(),
M_RT = round(mean(RT_ms), 1),
SD_RT = round(sd(RT_ms), 1),
Accuracy = round(mean(Accurate) * 100, 1),
.groups = "drop"
) |>
flextable() |>
flextable::set_table_properties(width = .8, layout = "autofit") |>
flextable::theme_zebra() |>
flextable::fontsize(size = 12) |>
flextable::fontsize(size = 12, part = "header") |>
flextable::align_text_col(align = "center") |>
flextable::set_caption(caption = "Reaction times and accuracy by condition in the lexical decision task.") |>
flextable::border_outer()
```
### `arrange()` — Sort Rows {-}
```{r arrange}
# Sort by RT (ascending)
lex_data |>
dplyr::arrange(RT_ms) |>
head(5)
# Sort descending
lex_data |>
dplyr::arrange(desc(RT_ms)) |>
head(5)
```
### `rename()` and `relocate()` {-}
```{r rename}
# Rename columns
lex_data |>
dplyr::rename(ReactionTime = RT_ms, Correct = Accurate) |>
head(3)
```
### `count()` — Quick Frequency Tables {-}
```{r count}
# How many observations per condition?
lex_data |>
dplyr::count(Condition)
# Cross-tabulate condition and accuracy
lex_data |>
dplyr::count(Condition, Accurate)
```
### Handling Missing Values {-}
```{r missing}
# Check for missing values
sum(is.na(lex_data$RT_ms))
colSums(is.na(lex_data))
# Remove rows with any missing value
lex_data_clean <- lex_data |>
tidyr::drop_na()
# Replace NA with a value (e.g., mean imputation — use cautiously!)
lex_data |>
dplyr::mutate(RT_ms = ifelse(is.na(RT_ms), mean(RT_ms, na.rm = TRUE), RT_ms))
```
---
::: {.callout-tip}
## Exercises: Working with Data
:::
**Q1. What does `dplyr::mutate()` do?**
```{r}
#| echo: false
#| label: "DAT_Q1"
check_question("It adds new columns or modifies existing columns, keeping all other columns and rows unchanged",
options = c(
"It adds new columns or modifies existing columns, keeping all other columns and rows unchanged",
"It removes rows that do not meet a condition",
"It summarises columns into a single value per group",
"It sorts the data frame by one or more columns"
),
type = "radio",
q_id = "DAT_Q1",
random_answer_order = TRUE,
button_label = "Check answer",
right = "Correct! mutate() transforms the data frame by computing new columns (or overwriting existing ones) without changing the number of rows. For example, mutate(RT_s = RT_ms / 1000) adds a new column RT_s that is the RT in seconds. The other options describe filter() (rows), summarise() (aggregation), and arrange() (sorting).",
wrong = "Think about what 'mutate' means: to change or transform. Which operation changes the columns of a data frame?")
```
---
**Q2. You want the mean RT for each participant across all conditions. Which dplyr pipeline is correct?**
```{r}
#| echo: false
#| label: "DAT_Q2"
check_question("lex_data |> group_by(Participant) |> summarise(M_RT = mean(RT_ms))",
options = c(
"lex_data |> group_by(Participant) |> summarise(M_RT = mean(RT_ms))",
"lex_data |> summarise(M_RT = mean(RT_ms)) |> group_by(Participant)",
"lex_data |> filter(Participant) |> mutate(M_RT = mean(RT_ms))",
"lex_data |> group_by(Participant) |> mutate(M_RT = mean(RT_ms))"
),
type = "radio",
q_id = "DAT_Q2",
random_answer_order = TRUE,
button_label = "Check answer",
right = "Correct! group_by() must come before summarise() — it tells R to apply the summary function separately within each group. The option with group_by() and mutate() is subtly different: mutate() would add a new column with each participant's mean RT to every row (without collapsing rows), whereas summarise() collapses to one row per participant. Both are useful but answer different questions.",
wrong = "The order of operations matters: which function defines the groups, and which computes the summary? Can you summarise before you have defined the groups?")
```
---
# Basic Visualisation with ggplot2 {#viz}
::: {.callout-note}
## Section Overview
**What you'll learn:** How to create basic plots using `ggplot2`; the layered grammar of graphics
**Key concept:** Every ggplot2 plot is built by adding layers — data, aesthetics, geometries, and themes
:::
`ggplot2` is R's most powerful and widely used plotting package. It is based on the **Grammar of Graphics**: the idea that every plot can be described by a consistent set of components.
## The Grammar of Graphics {-}
Every `ggplot2` plot has at least three components:
1. **Data**: the data frame containing your variables
2. **Aesthetics** (`aes()`): which variables map to which visual properties (x axis, y axis, colour, size, shape)
3. **Geometry** (`geom_*()`): how the data are visually represented (points, bars, lines, boxes)
Additional optional components include scales, facets, themes, and labels.
```r
ggplot(data = my_data, aes(x = variable1, y = variable2)) +
geom_point() +
theme_bw() +
labs(title = "My plot", x = "X label", y = "Y label")
```
## Histograms {-}
```{r hist, message=FALSE, warning=FALSE}
ggplot(lex_data, aes(x = RT_ms, fill = Condition)) +
geom_histogram(bins = 20, color = "white", alpha = 0.7) +
facet_wrap(~ Condition, ncol = 1) +
scale_fill_manual(values = c("steelblue", "tomato", "seagreen")) +
theme_bw() +
theme(legend.position = "none", panel.grid.minor = element_blank()) +
labs(title = "Distribution of reaction times by condition",
x = "Reaction time (ms)", y = "Count")
```
## Boxplots {-}
```{r boxplot, message=FALSE, warning=FALSE}
ggplot(lex_data, aes(x = Condition, y = RT_ms, fill = Condition)) +
geom_boxplot(alpha = 0.7, outlier.color = "gray40") +
stat_summary(fun = mean, geom = "point",
shape = 18, size = 3, color = "black") +
scale_fill_manual(values = c("steelblue", "tomato", "seagreen")) +
theme_bw() +
theme(legend.position = "none", panel.grid.minor = element_blank()) +
labs(title = "Reaction times by condition",
subtitle = "Diamond = group mean; box = median and IQR",
x = "Condition", y = "Reaction time (ms)")
```
## Bar Charts {-}
```{r barplot, message=FALSE, warning=FALSE}
lex_data |>
dplyr::group_by(Condition) |>
dplyr::summarise(M_RT = mean(RT_ms),
SE = sd(RT_ms) / sqrt(n()),
.groups = "drop") |>
ggplot(aes(x = Condition, y = M_RT, fill = Condition)) +
geom_col(alpha = 0.8, width = 0.6) +
geom_errorbar(aes(ymin = M_RT - SE, ymax = M_RT + SE),
width = 0.2, linewidth = 0.8) +
scale_fill_manual(values = c("steelblue", "tomato", "seagreen")) +
theme_bw() +
theme(legend.position = "none", panel.grid.minor = element_blank()) +
labs(title = "Mean reaction time by condition",
subtitle = "Error bars = ±1 SE",
x = "Condition", y = "Mean RT (ms)")
```
## Scatter Plots {-}
```{r scatter, message=FALSE, warning=FALSE}
ggplot(lex_data, aes(x = Participant, y = RT_ms, color = Condition)) +
geom_point(alpha = 0.7, size = 2) +
scale_color_manual(values = c("steelblue", "tomato", "seagreen")) +
theme_bw() +
theme(panel.grid.minor = element_blank()) +
labs(title = "Individual RT observations by participant and condition",
x = "Participant ID", y = "Reaction time (ms)",
color = "Condition")
```
## Saving Plots {-}
```{r save_plot, eval=FALSE}
# Save the most recently displayed plot
ggsave(
filename = here::here("images", "my_plot.png"),
width = 8,
height = 5,
dpi = 300
)
# Save a named plot object
my_plot <- ggplot(lex_data, aes(x = RT_ms)) + geom_histogram()
ggsave(
plot = my_plot,
filename = here::here("images", "histogram.pdf"),
width = 6,
height = 4
)
```
::: {.callout-tip}
## ggplot2 Quick Tips
- Add `theme_bw()` for a clean white background (LADAL standard)
- Add `theme(panel.grid.minor = element_blank())` to remove minor gridlines
- Use `scale_color_manual()` / `scale_fill_manual()` to control colours
- Use `facet_wrap(~ variable)` to create small multiples
- Use `labs()` to set title, subtitle, and axis labels
- Use `+ coord_flip()` to swap x and y axes (useful for long category names)
:::
---
::: {.callout-tip}
## Exercises: Visualisation
:::
**Q1. In ggplot2, what does `aes()` control?**
```{r}
#| echo: false
#| label: "VIZ_Q1"
check_question("The mapping between variables in the data and visual properties of the plot (axes, colour, shape, size)",
options = c(
"The mapping between variables in the data and visual properties of the plot (axes, colour, shape, size)",
"The type of plot geometry (e.g., histogram, boxplot, scatter)",
"The overall visual theme and background style",
"The axis labels and plot title"
),
type = "radio",
q_id = "VIZ_Q1",
random_answer_order = TRUE,
button_label = "Check answer",
right = "Correct! aes() stands for 'aesthetics' and specifies the mapping from data variables to visual properties: aes(x = RT, y = Accuracy) maps RT to the horizontal axis and Accuracy to the vertical axis; aes(color = Condition) maps the Condition variable to point/line colour. The geometry (what shape the data takes) is controlled by geom_*() functions. The theme controls non-data elements like background colour and grid lines. Labels are added with labs().",
wrong = "Aesthetics in ggplot2 are specifically about how data variables are translated into visual properties. Which of these options describes that mapping?")
```
---
**Q2. Which `geom_*()` function would you use to create a histogram?**
```{r}
#| echo: false
#| label: "VIZ_Q2"
check_question("geom_histogram()",
options = c("geom_histogram()", "geom_bar()", "geom_col()", "geom_density()"),
type = "radio",
q_id = "VIZ_Q2",
random_answer_order = FALSE,
button_label = "Check answer",
right = "Correct! geom_histogram() bins a continuous variable and displays the frequency of observations in each bin. geom_bar() counts the occurrences of a categorical variable (or uses stat = 'identity' for pre-computed counts). geom_col() plots a bar chart where the bar height is already a column in the data. geom_density() draws a smooth kernel density estimate rather than binned bars.",
wrong = "Think about what a histogram does: it shows the distribution of a *continuous* variable by dividing it into bins and counting observations in each. Which geom specifically does this?")
```
---
# Getting Help {#help}
::: {.callout-note}
## Section Overview
**What you'll learn:** How to find help efficiently when you are stuck — both within R and online
:::
Every R user gets stuck regularly. Knowing where to look for help is as important as knowing R itself.
## Help Within R {-}
```{r help_r, eval=FALSE}
# Help page for a specific function
?mean
help(mean)
# Search for functions related to a keyword
??regression
apropos("filter")
# See a function's arguments
args(ggplot)
# See examples of a function in action
example(boxplot)
```
RStudio's **Help** tab (bottom right pane) renders help pages with formatted descriptions, argument lists, and examples.
## Vignettes {-}
Many packages include **vignettes** — detailed guides that show how to use the package end-to-end. These are often more useful than the function-level help pages:
```{r vignettes, eval=FALSE}
# List all vignettes for a package
vignette(package = "dplyr")
# Open a specific vignette
vignette("dplyr")
vignette("ggplot2-specs")
```
## Reading Error Messages {-}
Error messages are your friend — they tell you exactly what went wrong. Common error patterns:
::: {.callout-warning}
## Common Errors and What They Mean
**`object 'x' not found`**
→ The object `x` does not exist in your environment. Did you run the line that creates it? Is it spelled correctly (case-sensitive)?
**`could not find function "ggplot"`**
→ The package containing this function is not loaded. Did you run `library(ggplot2)`?
**`Error in read.csv("data.csv") : cannot open file`**
→ R cannot find the file. Check your working directory (`getwd()`), use `here::here()`, and check for typos in the filename.
**`non-numeric argument to binary operator`**
→ You tried to do arithmetic on a character string. Check the type of your object with `class()`.
**`NAs introduced by coercion`**
→ R tried to convert a character to numeric but could not. The unconvertible values became `NA`. Inspect the affected column for unexpected text.
**`object of type 'closure' is not subsettable`**
→ You tried to index a function as if it were a data frame (e.g., `mean[1]`). Check whether you forgot parentheses somewhere.
:::
## Searching Online {-}
The R community is enormous and helpful. When you encounter an error:
1. **Copy the exact error message** and paste it into Google with "R" at the start
2. **Stack Overflow** ([stackoverflow.com](https://stackoverflow.com/questions/tagged/r)) has answers to most common R questions
3. **RStudio Community** ([community.rstudio.com](https://community.rstudio.com/)) is welcoming to beginners
4. **CRAN package pages** list vignettes, reference manuals, and NEWS files
5. **Package websites** (e.g., [dplyr.tidyverse.org](https://dplyr.tidyverse.org/)) have well-structured guides
::: {.callout-tip}
## Writing a Good Question
If you need to ask for help, always provide:
- A **minimal reproducible example** — the smallest piece of code that demonstrates the problem
- Your **session info**: `sessionInfo()`
- The **exact error message** (copy-paste, do not retype)
- What you **expected** to happen vs. what actually happened
The `reprex` package helps format reproducible examples: `install.packages("reprex")`
:::
## Key Online Resources {-}
```{r resources_table, echo=FALSE, message=FALSE, warning=FALSE}
data.frame(
Resource = c(
"R for Data Science",
"RStudio Cheatsheets",
"CRAN Task Views",
"Stack Overflow [r]",
"Tidyverse documentation",
"ggplot2 documentation",
"R Graph Gallery"
),
URL = c(
"r4ds.hadley.nz",
"posit.co/resources/cheatsheets",
"cran.r-project.org/web/views",
"stackoverflow.com/questions/tagged/r",
"tidyverse.org",
"ggplot2.tidyverse.org",
"r-graph-gallery.com"
),
Why_useful = c(
"Free online book; the best comprehensive introduction to R and the tidyverse",
"One-page quick references for popular packages (dplyr, ggplot2, RMarkdown, etc.)",
"Curated lists of R packages by topic (linguistics, NLP, spatial, etc.)",
"Answers to nearly every R question; search before posting",
"Official documentation for dplyr, ggplot2, tidyr, readr, and more",
"Function reference, articles, and extension gallery",
"Hundreds of example plots with full reproducible code"
)
) |>
dplyr::rename("Why useful" = Why_useful) |>
flextable() |>
flextable::set_table_properties(width = .99, layout = "autofit") |>
flextable::theme_zebra() |>
flextable::fontsize(size = 11) |>
flextable::fontsize(size = 11, part = "header") |>
flextable::align_text_col(align = "left") |>
flextable::set_caption(caption = "Key online resources for learning R.") |>
flextable::border_outer()
```
---
# Best Practices {#bestpractice}
::: {.callout-note}
## Section Overview
**What you'll learn:** Habits and conventions that make your R code more readable, reproducible, and robust
:::
Good coding habits matter more the longer your projects become. These practices are worth building from day one.
## Code Style {-}
- **Comment your code** liberally: `# This filters to English speakers only`
- Use **consistent naming**: `word_count` not `WordCount` or `wc`
- Keep **lines under 80 characters** (use line breaks inside functions)
- Add **spaces around operators**: `x <- 5 * (3 + 2)` not `x<-5*(3+2)`
- Load all packages at the **top of the script**
- Set the random seed at the top when using random processes: `set.seed(42)`
## Project Structure {-}
- Always work inside an **R Project** (`.Rproj`)
- Use `here::here()` for all file paths — never hardcode absolute paths like `"C:/Users/Martin/..."`
- Keep raw data **read-only** — never overwrite original files; save processed versions separately
- Use **version control** (Git) for anything important
## Reproducibility {-}
- Write all analyses in **R Notebooks or scripts** — never rely on Console-only work
- Render your notebook from scratch periodically to confirm it runs end-to-end
- End every notebook with `sessionInfo()` to record package versions
- Consider using `renv` to snapshot your package environment
## Environment Hygiene {-}
```{r hygiene, eval=FALSE}
# See all objects in your environment
ls()
# Remove a specific object
rm(my_temp_variable)
# Remove everything (use with caution!)
rm(list = ls())
# Check working directory
getwd()
# Change working directory (prefer R Projects over setwd())
setwd("path/to/folder") # avoid this; use R Projects instead
```
# Citation & Session Info {-}
::: {.callout-note}
## Citation
```{r citation-callout, echo=FALSE, results='asis'}
cat(
params$author, ". ",
params$year, ". *",
params$title, "*. ",
params$institution, ". ",
"url: ", params$url, " ",
"(Version ", params$version, "), ",
"doi: ", params$doi, ".",
sep = ""
)
```
```{r citation-bibtex, echo=FALSE, results='asis'}
key <- paste0(
tolower(gsub(" ", "", gsub(",.*", "", params$author))),
params$year,
tolower(gsub("[^a-zA-Z]", "", strsplit(params$title, " ")[[1]][1]))
)
cat("```\n")
cat("@manual{", key, ",\n", sep = "")
cat(" author = {", params$author, "},\n", sep = "")
cat(" title = {", params$title, "},\n", sep = "")
cat(" year = {", params$year, "},\n", sep = "")
cat(" note = {", params$url, "},\n", sep = "")
cat(" organization = {", params$institution, "},\n", sep = "")
cat(" edition = {", params$version, "}\n", sep = "")
cat(" doi = {", params$doi, "}\n", sep = "")
cat("}\n```\n")
```
:::
```{r fin}
sessionInfo()
```
::: {.callout-note}
## AI Transparency Statement
This tutorial was written with the assistance of **Claude** (claude.ai), a large language model created by Anthropic. Claude was used to draft and structure the entire tutorial, including all R code, conceptual explanations, and exercises. All content was reviewed and approved by Martin Schweinberger, who takes full responsibility for its accuracy.
:::
[Back to top](#intro)
[Back to HOME](/index.html)
# References {-}